-
Computed_GO_Component_IDs:
-
Computed_GO_Components:
-
Computed_GO_Function_IDs:
-
Computed_GO_Functions:
-
Computed_GO_Process_IDs:
-
Computed_GO_Processes:
-
Curated_GO_Component_IDs:
-
Curated_GO_Components:
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
No Results
-
Fasta :-
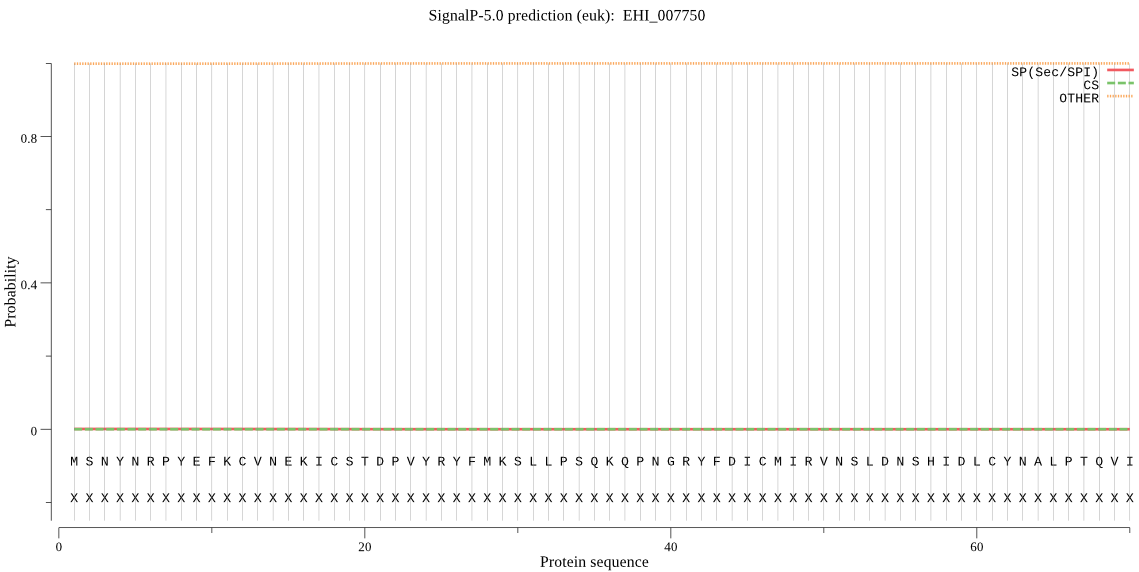
>EHI_007750
MSNYNRPYEFKCVNEKICSTDPVYRYFMKSLLPSQKQPNGRYFDICMIRVNSLDNSHIDL
CYNALPTQVIALYYTPTDSQLQKLTPPHKDYCGYYFFPYDIVEQQQSYFGGFNCIYYDID
WDDSSYLKNMVDGFDYQYVNKSTGTQNFDEVHFVASAPFINQRY
- Download Fasta
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/EHI_007750.fa
Sequence name : EHI_007750
Sequence length : 164
VALUES OF COMPUTED PARAMETERS
Coef20 : 3.682
CoefTot : -0.588
ChDiff : -5
ZoneTo : 8
KR : 1
DE : 0
CleavSite : 0
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 0.606 0.824 -0.007 0.415
MesoH : -0.849 -0.169 -0.444 0.145
MuHd_075 : 25.686 14.324 9.878 5.706
MuHd_095 : 8.251 11.321 2.258 2.151
MuHd_100 : 5.735 6.355 1.323 1.193
MuHd_105 : 17.117 7.804 3.315 3.855
Hmax_075 : 4.025 7.600 2.544 2.820
Hmax_095 : -8.663 1.488 -2.554 0.586
Hmax_100 : -9.800 -0.800 -2.462 -0.050
Hmax_105 : -0.467 3.383 -2.617 2.637
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.8798 0.1202
DFMC : 0.9239 0.0761
-
Fasta :-
>EHI_007750
ATGTCAAATTATAATAGACCATATGAATTTAAATGTGTTAATGAAAAGATTTGTTCAACT
GATCCAGTTTATAGATATTTCATGAAATCATTACTTCCATCTCAAAAACAACCAAATGGA
AGATATTTTGATATTTGTATGATTCGAGTTAATTCTCTTGATAATTCACATATTGATTTA
TGTTATAATGCTCTTCCAACTCAAGTTATTGCATTATATTATACTCCAACTGATTCACAA
TTACAAAAATTAACACCACCACATAAAGATTATTGTGGATATTATTTCTTCCCTTATGAT
ATCGTTGAACAACAACAATCCTATTTTGGTGGGTTTAATTGCATTTATTATGATATTGAT
TGGGATGATTCTAGTTATCTTAAAAATATGGTTGATGGTTTTGATTATCAATACGTTAAT
AAATCAACTGGAACACAGAATTTTGATGAAGTACATTTTGTTGCTTCAGCACCATTTATT
AATCAACGCTATTAA
- Download Fasta


