-
Computed_GO_Component_IDs:
-
Computed_GO_Components:
-
Computed_GO_Function_IDs:
-
Computed_GO_Functions:
-
Computed_GO_Process_IDs:
-
Computed_GO_Processes:
-
Curated_GO_Component_IDs:
-
Curated_GO_Components:
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
No Results
-
Fasta :-
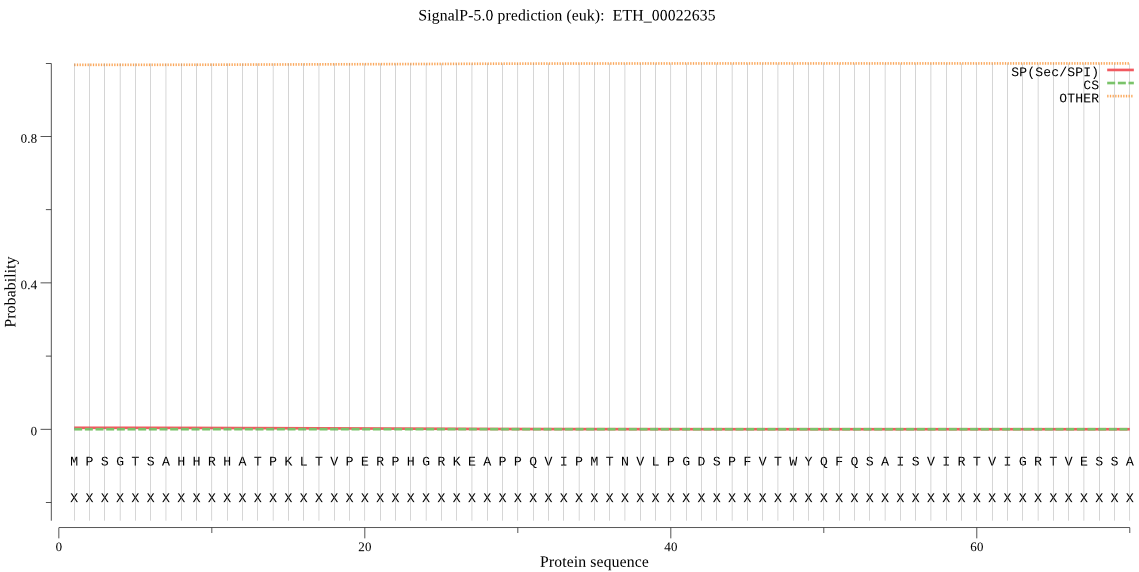
>ETH_00022635
MPSGTSAHHRHATPKLTVPERPHGRKEAPPQVIPMTNVLPGDSPFVTWYQFQSAISVIRT
VIGRTVESSATIPTLPEDSSTMQAFLLRIERRYTQMGLEPREWGNARIDHLVGPALTYWM
YLQRTIDLSDWATLQRRLFERFDKTMSQSKLLTELAKVRWSGNPKEYTYRFAAVAERGLG
VAPDELADYYCTGLPTDLHLLITNNGQVKYQSWEQAATAAARLYEPKQSRQARMPFQSPL
SEETNTSAQRKELLRAMGTRKGLSRFRKRGGSLGQHRSRVKRLRKRRRIAAGSLLRQQKT
AIGQTRQRLCHTGGGRLAPKSLATKEERCVVWERRRYFALSSQGVLVRPSCTLGLREVSL
VLKQSNDCS
- Download Fasta
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/ETH_00022635.fa
Sequence name : ETH_00022635
Sequence length : 369
VALUES OF COMPUTED PARAMETERS
Coef20 : 3.271
CoefTot : -1.076
ChDiff : 29
ZoneTo : 19
KR : 2
DE : 0
CleavSite : 12
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 0.712 1.041 0.051 0.431
MesoH : -0.532 0.058 -0.386 0.136
MuHd_075 : 24.844 13.131 5.918 5.131
MuHd_095 : 14.742 5.921 2.670 3.079
MuHd_100 : 9.106 4.396 2.568 2.579
MuHd_105 : 4.427 1.704 3.076 1.980
Hmax_075 : 5.800 2.400 -1.219 2.193
Hmax_095 : 7.175 0.525 -1.999 1.990
Hmax_100 : 7.000 1.300 -2.035 1.980
Hmax_105 : 3.733 2.000 -1.838 2.510
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.9390 0.0610
DFMC : 0.9067 0.0933
-
Fasta :-
>ETH_00022635
ATGCCGAGCGGGACGTCAGCCCATCATCGCCATGCGACTCCGAAGCTCACTGTGCCAGAG
CGGCCGCATGGGCGTAAGGAGGCGCCCCCGCAGGTGATACCTATGACCAATGTTCTGCCA
GGGGACAGTCCCTTCGTCACGTGGTACCAATTCCAGTCCGCTATAAGTGTGATAAGAACT
GTTATAGGGAGGACCGTGGAATCCAGCGCAACAATTCCCACACTACCAGAAGACAGCAGC
ACTATGCAGGCCTTTCTGCTGCGCATTGAGCGCCGGTACACCCAGATGGGATTGGAACCG
CGCGAGTGGGGAAATGCACGTATAGACCACCTTGTGGGCCCGGCTCTAACGTATTGGATG
TATCTACAGAGAACTATCGACCTAAGCGATTGGGCGACACTACAGCGCAGGCTTTTTGAG
CGGTTTGACAAAACAATGTCGCAGAGTAAATTGTTAACAGAGTTGGCTAAAGTACGGTGG
AGTGGTAACCCGAAAGAATACACGTACCGGTTTGCGGCTGTAGCGGAGCGTGGTTTGGGC
GTTGCCCCCGACGAACTCGCTGACTATTACTGTACCGGGCTACCGACGGACCTCCATCTT
CTAATAACCAATAACGGGCAAGTGAAATATCAGTCGTGGGAACAAGCGGCGACAGCCGCA
GCACGGCTCTACGAGCCCAAGCAGAGTCGACAGGCCCGAATGCCATTCCAGTCTCCCTTG
AGTGAGGAAACCAACACCTCAGCACAGCGGAAAGAACTTCTACGAGCGATGGGCACGCGG
AAGGGACTCAGCCGCTTTCGGAAACGCGGCGGGAGTCTGGGTCAGCATCGATCCCGAGTG
AAGAGACTCCGGAAGCGACGACGGATAGCAGCAGGAAGTCTGCTGAGGCAACAGAAGACG
GCTATAGGTCAGACGCGACAGAGATTGTGCCACACTGGTGGAGGGAGACTTGCACCAAAG
AGTCTTGCGACCAAGGAGGAGCGTTGTGTTGTGTGGGAGCGAAGGCGGTACTTCGCGCTG
AGCTCGCAGGGAGTCCTTGTGAGGCCCTCTTGTACACTGGGGCTTCGGGAAGTCTCATTA
GTCCTAAAACAGTCGAACGACTGCAGCTGA
- Download Fasta
|
ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method |
|
ETH_00022635 | 241 S | QSPLSEETN | 0.993 | unsp | ETH_00022635 | 241 S | QSPLSEETN | 0.993 | unsp | ETH_00022635 | 241 S | QSPLSEETN | 0.993 | unsp | ETH_00022635 | 272 S | KRGGSLGQH | 0.993 | unsp | ETH_00022635 | 13 T | HRHATPKLT | 0.995 | unsp | ETH_00022635 | 56 S | QSAISVIRT | 0.994 | unsp |


