-
Computed_GO_Component_IDs:
-
Computed_GO_Components:
-
Computed_GO_Function_IDs:
-
Computed_GO_Functions:
-
Computed_GO_Process_IDs:
-
Computed_GO_Processes:
-
Curated_GO_Component_IDs:
-
Curated_GO_Components:
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
No Results
-
Fasta :-
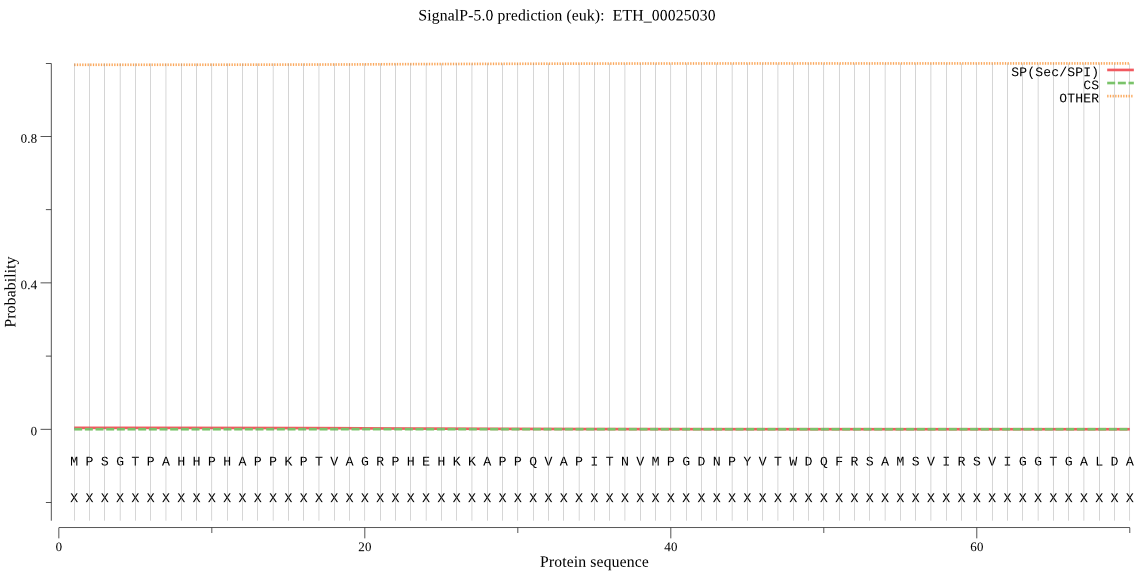
>ETH_00025030
MPSGTPAHHPHAPPKPTVAGRPHEHKKAPPQVAPITNVMPGDNPYVTWDQFRSAMSVIRS
VIGGTGALDATFPILREDGSNVQAVLLGVERRYIQMGMEPRDWGSALIDHLVGFALTY
- Download Fasta
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/ETH_00025030.fa
Sequence name : ETH_00025030
Sequence length : 118
VALUES OF COMPUTED PARAMETERS
Coef20 : 2.379
CoefTot : -4.637
ChDiff : 0
ZoneTo : 41
KR : 4
DE : 1
CleavSite : 0
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 0.371 1.000 0.003 0.461
MesoH : -0.746 0.220 -0.395 0.168
MuHd_075 : 24.010 20.119 6.863 6.741
MuHd_095 : 14.093 13.451 5.118 3.139
MuHd_100 : 22.458 18.387 6.998 4.782
MuHd_105 : 26.910 22.451 8.423 5.787
Hmax_075 : -1.200 7.100 -3.097 2.920
Hmax_095 : 0.700 5.000 -3.763 2.354
Hmax_100 : 8.700 9.600 -0.628 3.430
Hmax_105 : 12.017 15.633 0.731 4.580
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.9902 0.0098
DFMC : 0.9930 0.0070
-
Fasta :-
>ETH_00025030
ATGCCCAGCGGGACACCGGCTCATCATCCTCATGCTCCTCCGAAACCCACCGTGGCAGGG
CGGCCGCATGAACATAAGAAGGCGCCCCCACAGGTGGCACCTATAACCAATGTTATGCCA
GGGGACAATCCGTACGTCACGTGGGACCAATTCCGCTCCGCTATGAGTGTGATAAGAAGT
GTTATAGGCGGAACCGGGGCATTGGACGCAACGTTTCCCATACTTCGTGAAGACGGCAGC
AATGTGCAGGCCGTTTTGCTGGGCGTCGAACGACGGTACATCCAGATGGGAATGGAGCCC
CGCGACTGGGGGAGCGCACTTATTGACCATCTTGTGGGCTTCGCTCTAACGTATTGA
- Download Fasta
|
ID | Site | Peptide | Score | Method |
|
ETH_00025030 | 56 S | RSAMSVIRS | 0.993 | unsp |


