-
Computed_GO_Component_IDs:
-
Computed_GO_Components:
-
Computed_GO_Function_IDs:
-
Computed_GO_Functions:
-
Computed_GO_Process_IDs:
-
Computed_GO_Processes:
-
Curated_GO_Component_IDs:
-
Curated_GO_Components:
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
No Results
-
Fasta :-
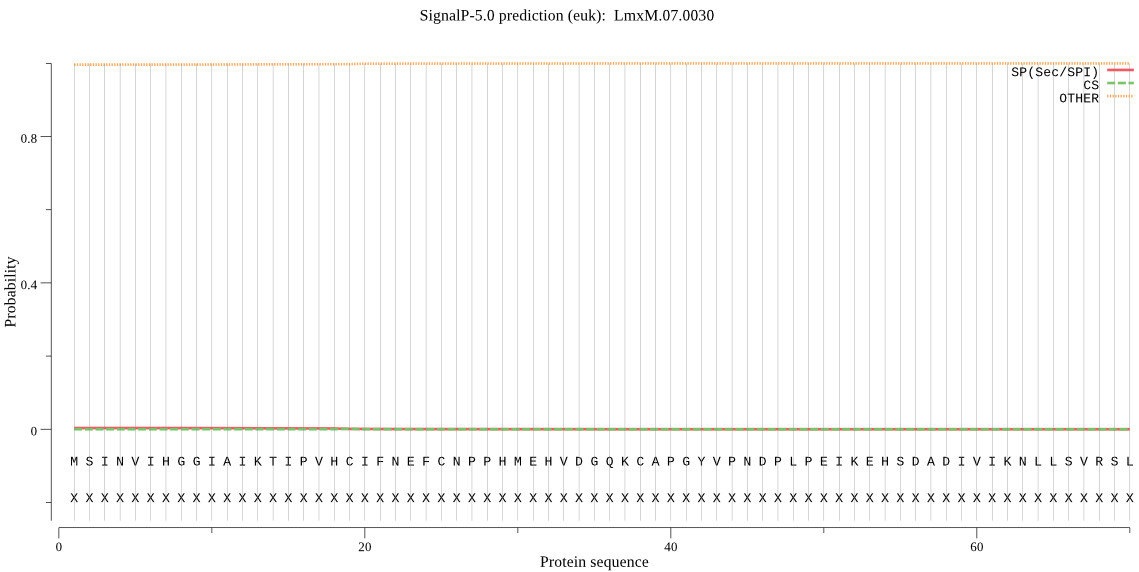
>LmxM.07.0030
MSINVIHGGIAIKTIPVHCIFNEFCNPPHMEHVDGQKCAPGYVPNDPLPEIKEHSDADIV
IKNLLSVRSLHDAEKNAFTALVRVMIGDRDTQTLGDNTEADQYALIGVPVIVYGPGSIRV
ANQENEFVPKKHVDDCGKKRAIHPTRKNHDM
- Download Fasta
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/470
Sequence name : 470
Sequence length : 151
VALUES OF COMPUTED PARAMETERS
Coef20 : 3.749
CoefTot : -1.359
ChDiff : -4
ZoneTo : 22
KR : 1
DE : 0
CleavSite : 0
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 1.129 1.465 0.177 0.611
MesoH : -1.202 0.075 -0.532 0.140
MuHd_075 : 27.205 27.194 10.762 7.558
MuHd_095 : 8.177 11.379 5.474 2.859
MuHd_100 : 6.263 11.760 5.488 2.481
MuHd_105 : 10.383 13.077 6.426 2.954
Hmax_075 : 17.588 22.050 5.304 7.610
Hmax_095 : 14.700 18.200 5.149 6.190
Hmax_100 : 12.600 16.200 4.338 5.480
Hmax_105 : 13.400 17.383 5.587 6.008
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.8654 0.1346
DFMC : 0.9249 0.0751
-
Fasta :-
>LmxM.07.0030
ATGTCGATCAATGTGATTCACGGCGGTATCGCGATAAAGACGATTCCTGTCCATTGCATT
TTCAACGAGTTCTGCAACCCACCTCACATGGAACATGTAGATGGCCAAAAATGCGCTCCT
GGCTACGTCCCGAACGATCCGCTTCCGGAGATAAAGGAACACAGTGATGCTGACATAGTC
ATCAAAAACCTTCTCTCCGTCCGCTCGCTCCACGACGCCGAGAAGAATGCCTTCACTGCA
CTGGTTCGTGTTATGATTGGTGACCGGGACACGCAGACGCTTGGTGACAACACGGAGGCT
GACCAGTATGCTCTTATCGGCGTTCCTGTTATCGTCTACGGCCCTGGAAGCATCAGAGTT
GCGAACCAGGAGAATGAGTTTGTCCCGAAGAAGCATGTGGATGACTGTGGGAAGAAGCGG
GCCATTCACCCTACGCGCAAGAACCATGATATGTAG
- Download Fasta
|
ID | Site | Peptide | Score | Method |
|
LmxM.07.0030 | 55 S | IKEHSDADI | 0.991 | unsp |


