-
Computed_GO_Component_IDs:
-
Computed_GO_Components:
-
-
-
Computed_GO_Process_IDs: GO:0006508
-
Computed_GO_Processes: proteolysis
-
-
-
-
-
Curated_GO_Processes:
|
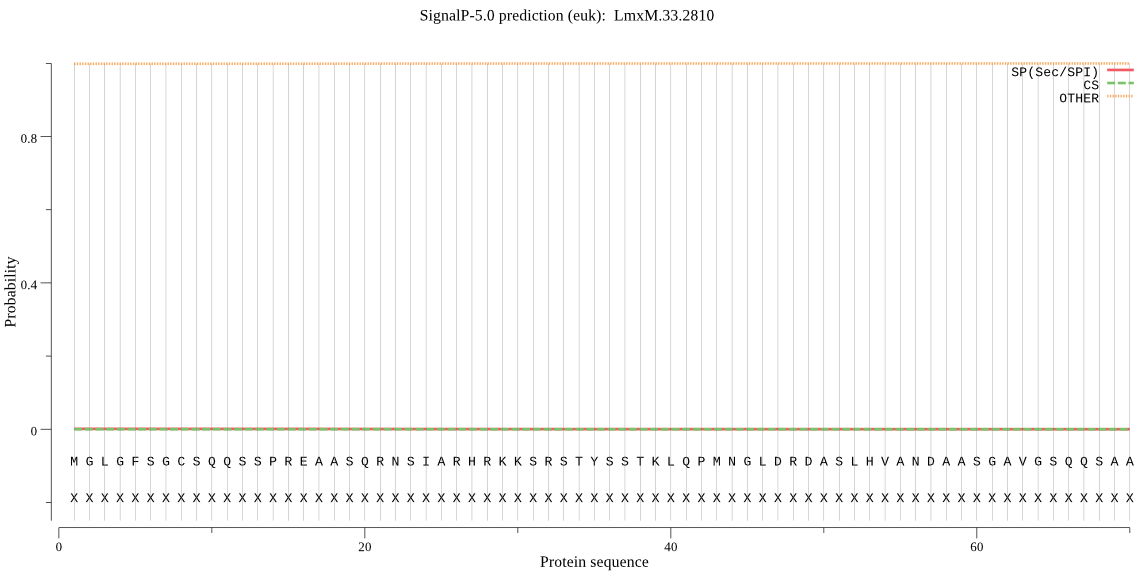
_ID | Prediction | OTHER | SP | mTP | CS_Position |
|
LmxM.33.2810 | OTHER | 0.999593 | 0.000032 | 0.000375 | |
No Results
-
Fasta :-
>LmxM.33.2810
MGLGFSGCSQQSSPREAASQRNSIARHRKKSRSTYSSTKLQPMNGLDRDASLHVANDAAS
GAVGSQQSAASLTVQGPEADDAHWVFQRPPNNQRTFYFQEDNLEFSSQFDSGNLIQVERV
GTFHYRMYTAMDCGNAPWQTNNRQWFHFSVRGGSKGVVVTICFVGMAHSNMFTYDWMPVM
AIVPTRPQYTRIAGKAHVEALEKMPETPGYPLLVHKPVSKDDADSDGGGDEGDNDADASA
AGANNNAGTGGIAFSISPTGKSSGTSKKKKAKKKSIAMNLTFNFKIETEIAVTYTPPQGR
PDSAAIYIASNHPYTYYTLQRNLGMWQEAARRSNALREMNVPVSDHQARRSSSEQRSGTD
SYALDGLPKVQPGSSGKSKKWVAGSNTGNAAAPYCAEFSEIYFHREVLCKSLEGRDVTLL
TISDCSSMTLERAPLLSKEDGLPHSSALGFTQRPYSFSGKQYVVLTARVHPGECPGSHLM
HGCIDFLMNCTDCRAAALRHNFVFCVVPMLNPDGVVRGHSRVDSNGVDLNRMYRDPSRKQ
HPAPYAVLALLRSLGSRVALFIDMHAHANKRGTFFYGNSMDWGNQIENLLYAKLVSLNTP
YLDFRSCNFSEANMFAVSKSGKRKDSSSRVVAFTEVGIVHGYTIETSHVMADTMNPVALL
SNHQADQLDTALLSPPPVMHSPATFHDTARAMLLGLLDLKGMNPISRLPLTQFHSTRGLA
LWLQRQLQIESAETLFAQAFKMYGKEAQASTGEFGNALLATIMRSMAAEEYPEKITIREA
RLLPRITFSGVRSFVPLETAIVLLSQTAPTGPPRSLLYGSGNCGAGSGTAAAGGTGCKGG
GSAQRRGTSPSSGNSCPTTVPPVVIATRRRSKPILLTSDTTVET
- Download Fasta
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/612
Sequence name : 612
Sequence length : 884
VALUES OF COMPUTED PARAMETERS
Coef20 : 3.642
CoefTot : -0.778
ChDiff : 16
ZoneTo : 46
KR : 8
DE : 1
CleavSite : 30
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 1.618 1.712 0.207 0.585
MesoH : 0.130 0.312 -0.110 0.247
MuHd_075 : 43.217 21.365 11.960 7.515
MuHd_095 : 23.665 14.395 6.072 5.394
MuHd_100 : 26.911 13.012 6.156 5.193
MuHd_105 : 35.236 14.488 8.699 6.684
Hmax_075 : 4.433 7.117 2.639 1.867
Hmax_095 : 3.000 -2.300 -3.628 -0.330
Hmax_100 : 3.000 -0.700 -4.177 0.030
Hmax_105 : 4.317 -1.167 -1.892 0.630
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.7600 0.2400
DFMC : 0.3334 0.6666
This protein is probably imported in mitochondria.
f(Ser) = 0.2174 f(Arg) = 0.1087 CMi = 0.97847
CMi is the Chloroplast/Mitochondria Index
It has been proposed by Von Heijne et al
(Eur J Biochem,1989, 180: 535-545)
-
Fasta :-
>LmxM.33.2810
ATGGGGCTGGGCTTCAGCGGCTGCTCCCAGCAGAGTTCGCCTAGGGAGGCCGCCTCGCAG
CGGAACTCGATAGCGAGGCACCGCAAAAAATCTAGATCCACTTATAGCAGCACGAAGTTG
CAGCCGATGAACGGGTTAGATCGAGATGCTTCTCTGCACGTCGCTAATGACGCGGCAAGC
GGCGCTGTAGGATCGCAGCAAAGCGCTGCAAGCCTGACGGTGCAAGGGCCGGAGGCGGAT
GACGCGCACTGGGTGTTTCAGAGACCGCCGAACAACCAGCGCACATTCTACTTTCAGGAG
GACAATCTTGAGTTCTCCTCGCAGTTTGACAGTGGCAACTTGATTCAGGTGGAGCGGGTG
GGCACCTTCCACTACCGCATGTACACGGCGATGGACTGCGGCAACGCACCATGGCAGACC
AACAACCGCCAGTGGTTTCACTTCTCCGTCCGCGGCGGCAGCAAGGGCGTTGTCGTCACG
ATCTGCTTTGTCGGCATGGCACACAGCAATATGTTCACCTATGATTGGATGCCGGTCATG
GCAATTGTGCCTACGCGACCGCAGTACACTCGCATCGCTGGAAAAGCGCACGTGGAGGCC
CTCGAGAAAATGCCAGAGACTCCCGGTTATCCTCTGCTTGTGCACAAGCCCGTGTCAAAG
GACGACGCCGACAGTGACGGTGGCGGCGATGAAGGTGACAACGATGCCGATGCCAGCGCT
GCTGGCGCCAACAACAATGCGGGCACTGGCGGAATCGCTTTCTCCATCTCGCCTACGGGC
AAGTCCTCTGGGACCAGCAAGAAGAAAAAGGCAAAGAAGAAGAGCATTGCTATGAACCTC
ACCTTCAACTTCAAAATTGAGACAGAGATCGCGGTAACGTACACCCCGCCTCAGGGGCGC
CCAGACTCGGCCGCGATCTACATTGCAAGCAATCACCCTTATACCTACTACACACTGCAA
CGCAACCTTGGGATGTGGCAGGAGGCGGCGCGACGAAGTAACGCTTTGCGGGAGATGAAT
GTGCCGGTATCCGACCACCAAGCGCGGCGCTCCAGCTCCGAGCAGCGTAGCGGGACGGAC
TCGTACGCCCTAGACGGCCTGCCCAAAGTTCAGCCTGGAAGCAGCGGAAAGAGTAAGAAG
TGGGTTGCAGGTTCCAACACCGGAAACGCTGCTGCTCCATACTGTGCGGAGTTCAGCGAG
ATCTATTTCCACCGCGAGGTTCTCTGCAAATCGCTGGAGGGGCGCGACGTTACACTGCTG
ACGATTAGCGACTGCAGTTCGATGACGTTAGAGCGGGCACCCCTCCTTTCCAAGGAAGAC
GGCCTTCCCCACTCTTCAGCTCTCGGCTTTACGCAGAGGCCGTACTCCTTTTCTGGCAAG
CAGTACGTGGTGCTGACGGCTCGTGTGCATCCAGGCGAATGCCCAGGCAGCCATCTTATG
CACGGCTGTATTGATTTTCTCATGAACTGCACAGACTGTCGAGCGGCGGCGCTGCGTCAC
AACTTTGTCTTCTGCGTGGTTCCGATGCTGAACCCCGACGGCGTAGTTCGCGGCCACAGC
CGCGTCGACTCAAACGGCGTCGATCTCAACCGCATGTACAGGGACCCGTCACGTAAGCAA
CACCCCGCACCATACGCGGTTCTCGCACTTCTGCGGTCTCTAGGCAGCCGCGTGGCGCTT
TTCATCGACATGCACGCACATGCCAATAAGCGGGGCACCTTCTTTTACGGTAACAGCATG
GATTGGGGCAACCAGATAGAAAACCTGCTTTACGCAAAGCTGGTCTCGCTTAACACGCCC
TACCTCGACTTCCGCAGCTGCAACTTCTCGGAGGCGAACATGTTCGCCGTTAGCAAGTCT
GGCAAACGCAAGGACTCCTCCAGCCGCGTCGTGGCCTTTACCGAGGTCGGTATCGTGCAC
GGCTACACCATCGAGACCTCACACGTGATGGCTGACACAATGAATCCAGTCGCGCTGCTG
TCGAATCATCAAGCCGACCAACTGGATACTGCCCTTTTAAGCCCACCCCCTGTTATGCAC
TCACCAGCTACCTTTCACGACACTGCTCGTGCGATGCTGCTGGGACTGCTGGACCTTAAG
GGTATGAACCCCATAAGTCGTCTACCCTTGACACAGTTTCACTCCACCCGCGGACTCGCT
CTTTGGCTGCAGCGGCAGCTGCAGATTGAATCAGCGGAGACGCTTTTTGCGCAGGCCTTC
AAGATGTACGGCAAGGAAGCTCAGGCCTCCACAGGCGAGTTTGGGAACGCTCTGCTCGCC
ACCATCATGAGAAGCATGGCGGCAGAGGAGTATCCGGAGAAGATCACAATCAGGGAGGCG
CGGTTGCTGCCCCGAATCACCTTCAGCGGTGTTCGGAGCTTTGTGCCACTAGAGACAGCA
ATTGTGCTGCTGTCGCAAACGGCGCCAACGGGGCCCCCGCGCTCGCTTCTGTACGGCAGC
GGCAACTGTGGGGCGGGAAGCGGCACTGCCGCTGCCGGGGGAACCGGCTGCAAAGGCGGC
GGCAGCGCTCAGCGCCGTGGCACGTCTCCCTCGAGTGGGAACAGCTGCCCGACTACGGTC
CCGCCGGTGGTGATCGCGACGCGGCGCCGTTCGAAGCCAATTCTTCTCACCAGCGATACA
ACTGTTGAAACGTAG
- Download Fasta
- title: Zn binding site
- coordinates: H470,E473,H565
|
ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method |
|
LmxM.33.2810 | 33 S | KKSRSTYSS | 0.996 | unsp | LmxM.33.2810 | 33 S | KKSRSTYSS | 0.996 | unsp | LmxM.33.2810 | 33 S | KKSRSTYSS | 0.996 | unsp | LmxM.33.2810 | 37 S | STYSSTKLQ | 0.993 | unsp | LmxM.33.2810 | 219 S | HKPVSKDDA | 0.997 | unsp | LmxM.33.2810 | 225 S | DDADSDGGG | 0.994 | unsp | LmxM.33.2810 | 266 S | SSGTSKKKK | 0.994 | unsp | LmxM.33.2810 | 351 S | QARRSSSEQ | 0.997 | unsp | LmxM.33.2810 | 352 S | ARRSSSEQR | 0.995 | unsp | LmxM.33.2810 | 353 S | RRSSSEQRS | 0.992 | unsp | LmxM.33.2810 | 357 S | SEQRSGTDS | 0.997 | unsp | LmxM.33.2810 | 378 S | SSGKSKKWV | 0.991 | unsp | LmxM.33.2810 | 437 S | APLLSKEDG | 0.997 | unsp | LmxM.33.2810 | 520 S | VRGHSRVDS | 0.995 | unsp | LmxM.33.2810 | 537 S | YRDPSRKQH | 0.997 | unsp | LmxM.33.2810 | 620 S | AVSKSGKRK | 0.995 | unsp | LmxM.33.2810 | 626 S | KRKDSSSRV | 0.995 | unsp | LmxM.33.2810 | 13 S | SQQSSPREA | 0.996 | unsp | LmxM.33.2810 | 31 S | HRKKSRSTY | 0.993 | unsp |


