-
-
-
Computed_GO_Function_IDs: GO:0008233
-
-
Computed_GO_Process_IDs: GO:0006465
-
-
Curated_GO_Component_IDs:
-
Curated_GO_Components:
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
No Results
-
Fasta :-
>PKNH_0420200
MGLIRDIKNSLVHTYVCLRRNRMDFHGQKLAFLIKNIIFTISTVVSIAVGYYKEDLALSA
YIILAGTVLSALLIMPTWPIYNKNNIHWESANESMNDRKRR
- Download Fasta
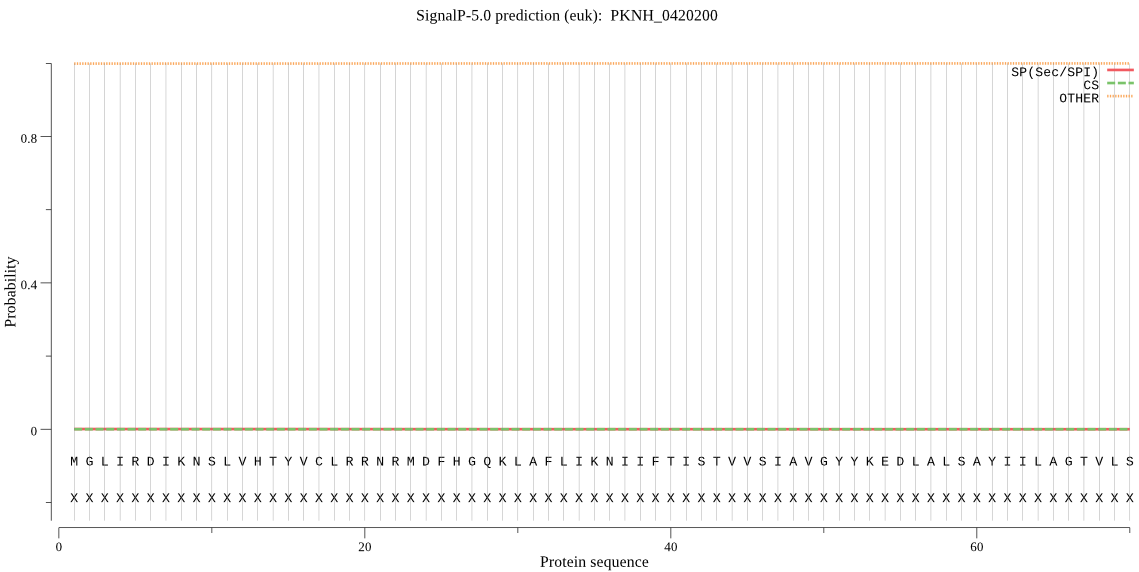
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/PKNH_0420200.fa
Sequence name : PKNH_0420200
Sequence length : 101
VALUES OF COMPUTED PARAMETERS
Coef20 : 4.444
CoefTot : -1.416
ChDiff : 6
ZoneTo : 53
KR : 8
DE : 2
CleavSite : 15
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 2.082 2.241 0.412 0.764
MesoH : 0.044 0.812 -0.217 0.356
MuHd_075 : 38.114 27.412 10.357 9.556
MuHd_095 : 36.057 26.346 9.189 8.323
MuHd_100 : 44.903 29.041 11.780 9.755
MuHd_105 : 39.130 25.566 10.944 7.846
Hmax_075 : 18.550 26.833 6.376 7.105
Hmax_095 : 4.100 25.375 1.518 3.570
Hmax_100 : 16.400 30.200 4.444 6.310
Hmax_105 : 17.500 28.200 5.729 6.113
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.5780 0.4220
DFMC : 0.6379 0.3621
-
Fasta :-
>PKNH_0420200
ATGGGCCTAATTCGCGATATTAAAAACAGTCTAGTGCACACCTATGTGTGTCTCCGGAGG
AATCGCATGGATTTCCACGGTCAGAAGCTCGCATTTTTAATCAAAAATATCATCTTCACC
ATCAGCACAGTTGTATCTATTGCAGTCGGCTATTACAAGGAGGACTTAGCCTTAAGTGCC
TACATCATTTTGGCTGGTACTGTCTTGTCTGCATTGCTGATCATGCCCACGTGGCCCATA
TACAACAAGAACAATATTCACTGGGAAAGTGCTAATGAGTCCATGAACGATAGGAAGAGA
AGATAA
- Download Fasta


