-
Computed_GO_Component_IDs:
-
Computed_GO_Components:
-
Computed_GO_Function_IDs:
-
Computed_GO_Functions:
-
Computed_GO_Process_IDs:
-
Computed_GO_Processes:
-
Curated_GO_Component_IDs:
-
Curated_GO_Components:
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
No Results
-
Fasta :-
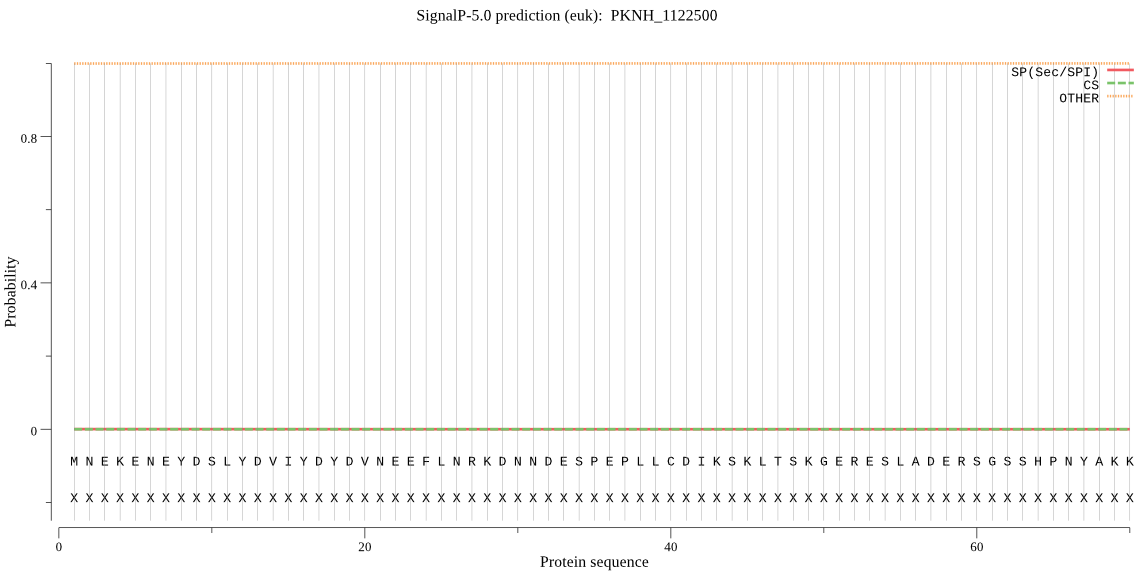
>PKNH_1122500
MNEKENEYDSLYDVIYDYDVNEEFLNRKDNNDESPEPLLCDIKSKLTSKGERESLADERS
GSSHPNYAKKNESEENSLQLSSIRLNDKNGDCSGEGQMKSTLTPQNNAPDSIKESEISSG
NVDPFKKREDLPFLKSSTQQNDSGNAEDGNSNAEAREEVKFVNPEKGGNHSPVVDAFTNS
TPKHTVGITNCENYNKEAPPHMRILSTDFQSTYSWQNVTVGKNNNNIVNRKNVLGESTPK
EQGTREECPQRDELKEGGAYHNNDVMIKGSGVKEKYKKLTFEDDIYKENADGKVEVQKEQ
GPNEDEKKKLEEKSDISEDKKNVEHSNDNNNSEEKNKTEINSNKNTTSENDGTSTFECNI
CFDDVRDPVVTKCGHLFCWLCLSAWIKKNNDCPVCKAEVSRENVIPLYGRGKNSTEHKYS
NKEEPRPTPKRKESVRRNNGYSNNVGLRASFGVWVNPFSFGMSYTNMTEEPYFYENRGGD
NRTQAETYQAEAASSFFFFLGFFLSLYILFYSS
- Download Fasta
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/PKNH_1122500.fa
Sequence name : PKNH_1122500
Sequence length : 513
VALUES OF COMPUTED PARAMETERS
Coef20 : 2.567
CoefTot : -0.134
ChDiff : -22
ZoneTo : 2
KR : 0
DE : 0
CleavSite : 0
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 2.488 2.071 0.345 0.873
MesoH : -0.424 -0.083 -0.421 0.191
MuHd_075 : 32.354 18.843 7.739 4.305
MuHd_095 : 24.900 16.517 6.875 4.578
MuHd_100 : 29.388 19.340 7.554 5.462
MuHd_105 : 28.821 19.188 7.032 5.573
Hmax_075 : -3.617 3.267 -3.174 1.680
Hmax_095 : -1.600 3.400 -2.424 2.420
Hmax_100 : -1.600 2.500 -2.424 2.420
Hmax_105 : -5.687 3.850 -3.289 2.205
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.9868 0.0132
DFMC : 0.9787 0.0213
-
Fasta :-
>PKNH_1122500
ATGAATGAGAAAGAAAACGAATATGATTCGCTGTACGATGTAATATACGATTATGATGTT
AACGAAGAATTCCTCAATAGGAAGGACAACAACGATGAGTCTCCAGAACCGCTGCTCTGT
GACATAAAAAGTAAATTAACTTCCAAGGGAGAAAGAGAAAGCTTGGCGGATGAGCGTTCA
GGAAGTTCCCACCCAAATTACGCAAAAAAAAATGAAAGCGAAGAAAACTCTTTACAATTA
TCGTCCATACGGTTAAACGATAAAAATGGCGATTGTTCAGGGGAAGGTCAAATGAAGAGT
ACGTTGACGCCTCAGAACAACGCCCCAGACAGTATCAAAGAGAGCGAAATAAGCTCCGGT
AACGTGGACCCTTTTAAAAAAAGGGAGGATCTGCCATTCTTAAAAAGTAGCACACAACAG
AATGATAGTGGAAACGCCGAGGATGGTAATTCCAATGCGGAAGCAAGAGAGGAGGTCAAA
TTTGTGAACCCCGAAAAGGGGGGAAATCACTCCCCAGTTGTTGACGCATTTACAAATAGC
ACCCCTAAACATACAGTGGGAATAACCAACTGCGAAAACTACAATAAAGAAGCGCCTCCC
CATATGAGGATACTATCAACCGATTTCCAGAGTACATACAGTTGGCAAAACGTAACTGTA
GGCAAAAATAATAACAACATAGTGAATCGTAAAAACGTATTAGGAGAAAGTACCCCCAAG
GAACAGGGTACCAGAGAGGAGTGTCCTCAGAGGGATGAGTTAAAAGAGGGGGGGGCATAT
CACAATAACGACGTTATGATAAAAGGTAGTGGGGTAAAGGAAAAATACAAAAAGCTTACC
TTCGAGGATGATATATATAAAGAGAATGCAGATGGAAAAGTTGAGGTACAAAAGGAACAA
GGCCCAAATGAGGATGAAAAAAAAAAATTGGAAGAAAAAAGTGACATTAGTGAAGACAAG
AAAAATGTAGAACACAGTAATGATAATAACAATTCGGAAGAAAAAAACAAAACGGAAATA
AATTCCAACAAGAATACAACATCAGAAAATGATGGTACAAGTACATTCGAATGTAACATT
TGTTTTGATGATGTGAGAGACCCAGTGGTAACGAAATGTGGCCATCTATTCTGTTGGTTA
TGTCTCTCCGCTTGGATTAAAAAAAATAATGACTGTCCTGTGTGCAAAGCTGAAGTTTCA
AGGGAAAATGTAATACCTCTGTATGGAAGGGGAAAAAATAGCACAGAACATAAATATTCG
AATAAGGAAGAACCTAGACCCACTCCGAAAAGAAAAGAAAGTGTTAGGAGAAATAATGGT
TATTCTAACAATGTAGGGTTAAGGGCTTCCTTTGGCGTTTGGGTGAACCCCTTTTCCTTT
GGAATGTCCTACACCAACATGACGGAAGAGCCGTACTTTTATGAAAACCGTGGTGGTGAC
AACAGAACCCAGGCAGAAACGTATCAAGCGGAAGCTGCATCCTCCTTTTTCTTTTTCCTG
GGTTTTTTTTTGTCCCTCTATATTTTGTTCTACTCATCTTAG
- Download Fasta
- title: Zn binding site
- coordinates: C358,C361,C373,H375,C378,C381,C392,C395
|
ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method |
|
PKNH_1122500 | 54 S | GERESLADE | 0.997 | unsp | PKNH_1122500 | 54 S | GERESLADE | 0.997 | unsp | PKNH_1122500 | 54 S | GERESLADE | 0.997 | unsp | PKNH_1122500 | 73 S | KKNESEENS | 0.995 | unsp | PKNH_1122500 | 111 S | NAPDSIKES | 0.998 | unsp | PKNH_1122500 | 286 Y | EDDIYKENA | 0.991 | unsp | PKNH_1122500 | 420 S | EHKYSNKEE | 0.997 | unsp | PKNH_1122500 | 428 T | EPRPTPKRK | 0.994 | unsp | PKNH_1122500 | 434 S | KRKESVRRN | 0.996 | unsp | PKNH_1122500 | 34 S | NNDESPEPL | 0.992 | unsp | PKNH_1122500 | 48 S | SKLTSKGER | 0.995 | unsp |


