-
-
-
Computed_GO_Function_IDs:
-
Computed_GO_Functions:
-
-
-
Curated_GO_Component_IDs:
-
Curated_GO_Components:
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
No Results
-
Fasta :-
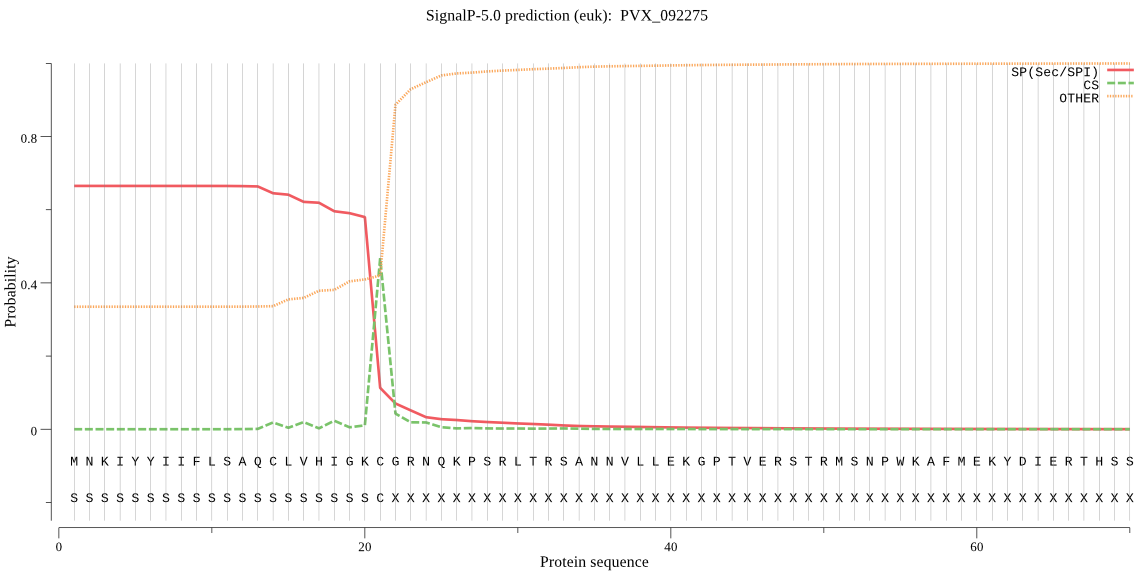
>PVX_092275
MNKIYYIIFLSAQCLVHIGKCGRNQKPSRLTRSANNVLLEKGPTVERSTRMSNPWKAFME
KYDIERTHSSGVRVDLGEDAEVENAKYRIPAGRCPVFGKGIVIENSDVSFLRPVATGDQK
LKDGGFAFPNANDHISPMTLANLKERYKDNVEMMKLNDIALCRTHAASFVMAGDQNSSYR
HPAVYDEKEKTCHMLYLSAQENMGPRYCSPDAQNRDAVFCFKPDKNESFENLVYLSKNVR
NDWDKKCPRKNLGNAKFGLWVDGNCEEIPYVKEVEAEDLRECNRIVFGASASDQPTQYEE
EMTDYQKIQQGFRQNNREMIKSAFLPVGAFNSDNFKSKGRGFNWANFDSVKKKCYIFNTK
PTCLINDKNFIATTALSHPQEVDLEFPCSIYKDEIEREIKKQSRNMNLYSVDGERIVLPR
IFISNDKESIKCPCEPERISNSTCNFYVCNCVEKRAEIKENNQVVIKEEFRDYYENGEEK
SNKQMLLIIIGITGGVCVVALASMAYFRKKANNDKYDKMDQAEGYGKPTTRKDEMLDPEA
SFWGEDKRASHTTPVLMEKPYY
- Download Fasta
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/PVX_092275.fa
Sequence name : PVX_092275
Sequence length : 562
VALUES OF COMPUTED PARAMETERS
Coef20 : 3.909
CoefTot : -0.858
ChDiff : 1
ZoneTo : 39
KR : 6
DE : 0
CleavSite : 0
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 2.341 2.724 0.627 0.891
MesoH : -1.249 -0.074 -0.574 0.063
MuHd_075 : 18.224 11.205 7.539 3.456
MuHd_095 : 23.684 20.043 9.781 5.526
MuHd_100 : 29.918 23.837 11.539 6.563
MuHd_105 : 35.585 21.386 11.402 7.797
Hmax_075 : 16.333 19.600 4.572 6.130
Hmax_095 : 15.400 22.500 6.218 -0.580
Hmax_100 : 15.700 23.500 6.218 6.080
Hmax_105 : 6.000 23.500 7.029 3.210
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.9001 0.0999
DFMC : 0.8788 0.1212
-
Fasta :-
>PVX_092275
ATGAATAAAATATACTACATAATCTTTTTAAGCGCCCAGTGCCTTGTGCACATTGGGAAG
TGCGGGCGAAACCAGAAGCCGAGCAGGCTGACCCGTAGCGCCAACAACGTTCTACTGGAA
AAGGGGCCTACCGTTGAGAGAAGCACACGAATGAGTAACCCCTGGAAAGCGTTCATGGAA
AAATACGACATCGAAAGAACACACAGTTCTGGGGTTCGAGTGGATTTAGGGGAAGATGCA
GAAGTGGAAAATGCAAAGTACAGAATTCCAGCTGGAAGATGTCCTGTTTTTGGAAAGGGT
ATCGTTATAGAGAATTCTGACGTTAGCTTCTTAAGACCTGTGGCTACAGGAGATCAGAAG
CTGAAGGATGGAGGTTTCGCCTTCCCCAATGCGAATGACCATATCTCCCCAATGACATTA
GCGAACCTTAAGGAAAGGTATAAAGACAATGTAGAGATGATGAAGTTAAACGATATAGCT
TTGTGCAGAACCCACGCAGCTAGCTTTGTCATGGCAGGGGATCAAAATTCGTCCTACAGA
CACCCAGCTGTATACGACGAAAAGGAAAAAACATGCCACATGTTGTATTTATCAGCGCAG
GAAAATATGGGTCCGAGGTACTGCAGCCCAGATGCACAAAATAGAGATGCCGTGTTCTGC
TTCAAGCCAGATAAAAATGAAAGCTTTGAAAACCTGGTGTATTTGAGCAAAAATGTGCGT
AATGATTGGGATAAAAAATGCCCCCGTAAAAATTTAGGAAACGCCAAGTTCGGATTATGG
GTGGATGGGAACTGCGAAGAAATTCCATACGTTAAAGAAGTGGAGGCAGAGGATCTGCGC
GAATGCAACCGAATCGTTTTCGGAGCGAGTGCCTCAGATCAACCAACTCAGTATGAAGAA
GAAATGACGGATTATCAAAAAATACAACAAGGGTTTAGACAAAACAACCGAGAGATGATT
AAAAGTGCCTTTCTTCCAGTGGGTGCATTCAACTCGGATAATTTCAAAAGTAAAGGAAGA
GGATTTAACTGGGCAAATTTCGATTCTGTAAAAAAGAAGTGTTACATTTTTAATACCAAA
CCGACTTGCCTCATTAATGACAAAAATTTTATTGCAACAACGGCGTTATCTCACCCACAA
GAAGTAGACCTGGAGTTCCCCTGCAGCATATATAAAGACGAAATTGAAAGAGAAATTAAG
AAACAATCGAGGAACATGAATCTGTACAGTGTTGATGGGGAACGCATTGTCCTGCCGAGG
ATATTTATCTCCAACGATAAGGAGAGTATCAAATGTCCCTGCGAACCTGAGCGCATTTCC
AACAGTACCTGCAACTTTTACGTTTGTAACTGTGTAGAGAAAAGGGCGGAAATTAAGGAA
AATAACCAAGTTGTTATAAAGGAAGAATTTAGGGATTATTACGAAAATGGGGAGGAAAAA
TCGAACAAGCAGATGCTACTAATCATTATCGGAATAACTGGTGGCGTGTGCGTCGTCGCG
CTGGCCTCTATGGCCTACTTCAGGAAGAAGGCTAACAATGATAAGTATGACAAGATGGAC
CAGGCAGAGGGGTACGGGAAGCCCACCACCAGGAAGGACGAGATGCTCGACCCCGAGGCC
TCCTTCTGGGGCGAGGACAAGCGGGCCTCCCACACCACGCCCGTGCTGATGGAGAAGCCG
TACTACTGA
- Download Fasta
|
ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method |
|
PVX_092275 | 550 S | DKRASHTTP | 0.991 | unsp | PVX_092275 | 550 S | DKRASHTTP | 0.991 | unsp | PVX_092275 | 550 S | DKRASHTTP | 0.991 | unsp | PVX_092275 | 290 S | VFGASASDQ | 0.994 | unsp | PVX_092275 | 429 S | NDKESIKCP | 0.995 | unsp |


