-
Computed_GO_Component_IDs:
-
Computed_GO_Components:
-
Computed_GO_Function_IDs: GO:0004222
-
-
Computed_GO_Process_IDs: GO:0006508
-
Computed_GO_Processes: proteolysis
-
Curated_GO_Component_IDs:
-
Curated_GO_Components:
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
No Results
-
Fasta :-
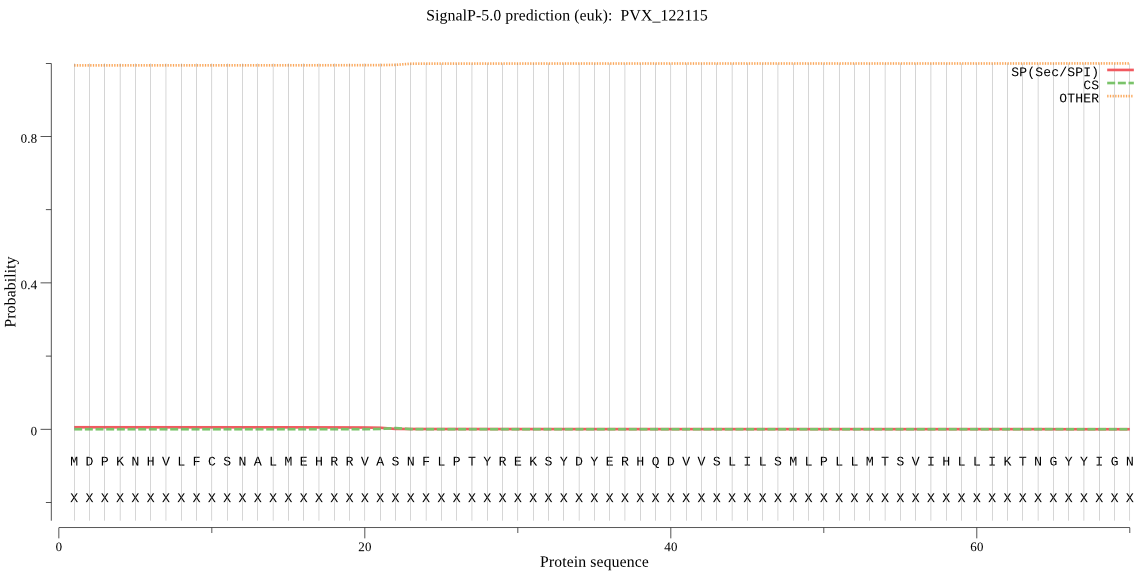
>PVX_122115
MDPKNHVLFCSNALMEHRRVASNFLPTYREKSYDYERHQDVVSLILSMLPLLMTSVIHLL
IKTNGYYIGNLFLMITYLFSFFFFILYYYNSYAVFVLFVFSSFFISLCLHEFAHALVAYK
YGDITMVYKGYLYLDIANYLDIFHTLIIPSITLFITGFGLPGNLYWIQLHFIRSRFQLSF
IYLSGPISDILCILLIVFFYNFYSYVRNSKNLSVQPHSILFISLATAASFLVDSFLLNIC
PIFGFDGWGIVEPYLPYCVNSLINEEIVNTYLSYICPLLVFIYFNFIETKYLFFTKIVNF
ILARILGIEISHVTIGVSAFPTLYAYLRNV
- Download Fasta
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/PVX_122115.fa
Sequence name : PVX_122115
Sequence length : 330
VALUES OF COMPUTED PARAMETERS
Coef20 : 3.876
CoefTot : -0.850
ChDiff : -1
ZoneTo : 29
KR : 4
DE : 2
CleavSite : 31
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 2.553 2.612 0.521 0.902
MesoH : 1.475 1.542 0.121 0.634
MuHd_075 : 21.296 21.688 7.837 5.696
MuHd_095 : 26.118 20.471 7.730 7.299
MuHd_100 : 29.591 21.997 8.842 7.828
MuHd_105 : 37.122 20.980 8.735 8.506
Hmax_075 : 7.612 12.483 1.125 2.660
Hmax_095 : 13.300 14.438 2.055 5.014
Hmax_100 : 8.800 13.700 1.922 4.500
Hmax_105 : 10.500 10.500 0.033 4.190
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.7607 0.2393
DFMC : 0.8705 0.1295
-
Fasta :-
>PVX_122115
ATGGATCCGAAAAACCACGTGCTCTTTTGCAGCAACGCACTGATGGAACACAGAAGAGTA
GCGAGCAATTTCCTTCCCACGTACAGAGAAAAAAGTTACGATTATGAGAGGCACCAAGAT
GTGGTATCGCTTATTCTGTCGATGCTCCCTTTATTAATGACAAGTGTAATTCACCTTTTG
ATAAAAACGAATGGTTACTATATTGGGAATTTATTCCTAATGATAACTTATTTATTTTCT
TTTTTTTTCTTCATTTTGTATTATTACAATTCTTACGCAGTGTTTGTCCTTTTTGTGTTT
TCCTCCTTTTTCATATCATTGTGTTTACACGAATTTGCCCACGCGCTTGTTGCGTATAAG
TACGGAGACATTACGATGGTTTACAAAGGCTACCTCTATCTGGACATTGCAAATTATTTA
GACATTTTCCACACGCTTATAATACCTTCTATAACGCTGTTTATAACCGGCTTTGGCCTA
CCAGGGAATTTATATTGGATACAGCTACACTTTATAAGAAGCAGATTTCAGCTGTCATTT
ATTTATTTATCGGGCCCCATATCAGACATTTTGTGTATCCTGCTAATTGTTTTTTTTTAC
AATTTTTACTCCTACGTTAGGAATAGCAAAAATTTAAGTGTGCAGCCGCACTCCATTTTA
TTCATATCCCTTGCCACTGCTGCTTCCTTCCTGGTGGACTCCTTTCTTTTAAACATTTGT
CCAATTTTTGGCTTTGACGGATGGGGGATCGTGGAGCCCTACCTCCCCTACTGTGTGAAT
AGCCTAATCAACGAAGAAATTGTGAACACCTATTTGTCCTACATATGCCCCCTCCTGGTT
TTTATATATTTTAACTTTATCGAAACCAAGTACCTCTTCTTCACAAAGATTGTTAACTTT
ATATTGGCGCGCATCCTCGGAATTGAAATTTCGCACGTGACAATTGGGGTCAGCGCGTTT
CCCACGTTGTATGCGTACCTCAGAAATGTGTAG
- Download Fasta
- title: active site
- coordinates: H110,E111,H114,N238,D246
|
ID | Site | Peptide | Score | Method |
|
PVX_122115 | 32 S | YREKSYDYE | 0.995 | unsp |


