-
Computed_GO_Component_IDs:
-
Computed_GO_Components:
-
Computed_GO_Function_IDs:
-
Computed_GO_Functions:
-
Computed_GO_Process_IDs:
-
Computed_GO_Processes:
-
Curated_GO_Component_IDs:
-
Curated_GO_Components:
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
No Results
-
Fasta :-
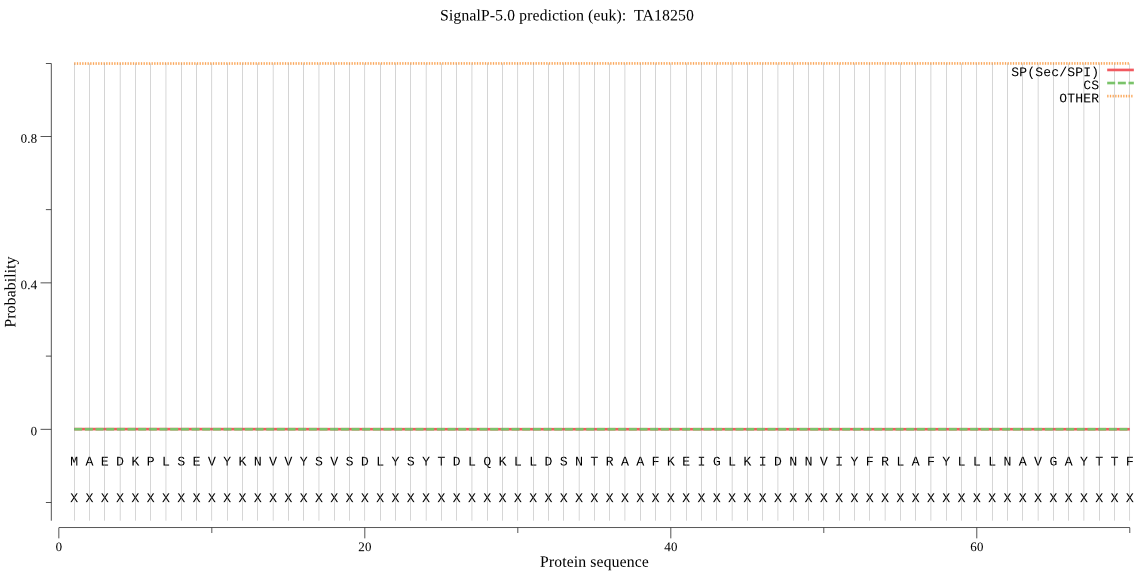
>TA18250
MAEDKPLSEVYKNVVYSVSDLYSYTDLQKLLDSNTRAAFKEIGLKIDNNVIYFRLAFYLL
LNAVGAYTTFFVHVEKSKFLLKVMLIVFFSLFSVYIAYDKVLLCRAHFVVKLKYGQELRG
WKELLKLVRYKRKGRKNRTETYTLPLGDLFFSDGECDYDHFVKLMEKLKPTLLSLDRKTK
KQD
- Download Fasta
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/TA18250.fa
Sequence name : TA18250
Sequence length : 183
VALUES OF COMPUTED PARAMETERS
Coef20 : 3.358
CoefTot : 0.141
ChDiff : 10
ZoneTo : 2
KR : 0
DE : 0
CleavSite : 0
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 1.882 2.371 0.258 0.773
MesoH : 0.022 1.082 -0.228 0.395
MuHd_075 : 23.814 11.984 5.852 4.039
MuHd_095 : 25.394 23.040 8.905 6.825
MuHd_100 : 21.086 20.080 7.759 5.682
MuHd_105 : 20.614 18.017 6.874 4.652
Hmax_075 : 2.567 10.500 -1.339 2.987
Hmax_095 : 4.300 15.800 0.679 5.020
Hmax_100 : 2.800 10.800 -0.022 3.740
Hmax_105 : 4.113 11.112 0.100 3.693
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.9111 0.0889
DFMC : 0.9356 0.0644
-
Fasta :-
>TA18250
ATGGCAGAGGATAAGCCCCTCAGCGAAGTGTATAAAAATGTAGTTTATTCTGTGTCGGAT
TTATACTCTTACACCGATCTCCAAAAGTTACTCGACTCAAATACGAGAGCAGCTTTTAAA
GAAATCGGTCTTAAAATCGATAATAATGTCATCTACTTCAGACTTGCTTTCTATTTACTT
CTGAACGCAGTCGGAGCCTACACCACCTTCTTCGTTCACGTAGAAAAGTCAAAATTTTTG
TTAAAGGTTATGTTGATTGTTTTTTTCTCTTTATTTTCCGTCTATATCGCCTACGATAAG
GTTCTCCTATGCAGAGCACATTTTGTCGTAAAACTCAAATATGGGCAAGAGTTAAGAGGT
TGGAAGGAACTTTTGAAATTAGTACGTTATAAAAGAAAGGGTCGAAAAAATAGGACTGAA
ACATACACATTACCCCTAGGTGATTTGTTTTTCTCAGACGGAGAGTGCGATTATGACCAT
TTTGTAAAATTAATGGAGAAGCTCAAACCGACGCTTCTTAGTCTTGACAGAAAAACTAAA
AAACAAGACTAA
- Download Fasta
|
ID | Site | Peptide | Score | Method |
|
TA18250 | 17 S | NVVYSVSDL | 0.993 | unsp |


