

| _ID | Prediction | OTHER | SP | mTP | CS_Position | |
|---|---|---|---|---|---|---|
| cgd3_2160 | OTHER | 0.999698 | 0.000122 | 0.000180 |
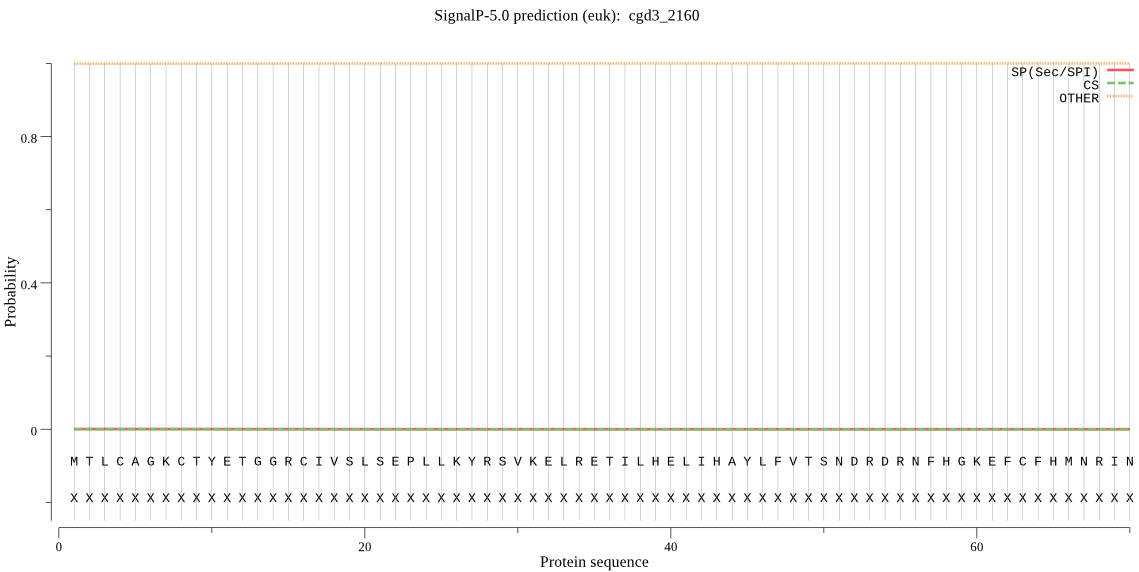
MTLCAGKCTYETGGRCIVSLSEPLLKYRSVKELRETILHELIHAYLFVTSNDRDRNFHGK EFCFHMNRINKMSGLNITIYHNFHDELNYYRRYIWRCDGVCRNHPPYYGYVRRSINRKPG PADSWWNFHRKTCGGCFVKEAKTLPQINNLESLGTRPNGQTSSPNMHQDDILEIIEISD
| ID | Site | Peptide | Score | Method | |
|---|---|---|---|---|---|
| cgd3_2160 | 29 S | LKYRSVKEL | 0.996 | unsp |