-
Computed_GO_Component_IDs: GO:0016021
-
-
Computed_GO_Function_IDs: GO:0004252
-
-
Computed_GO_Process_IDs:
-
Computed_GO_Processes:
-
Curated_GO_Component_IDs:
-
Curated_GO_Components:
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
|
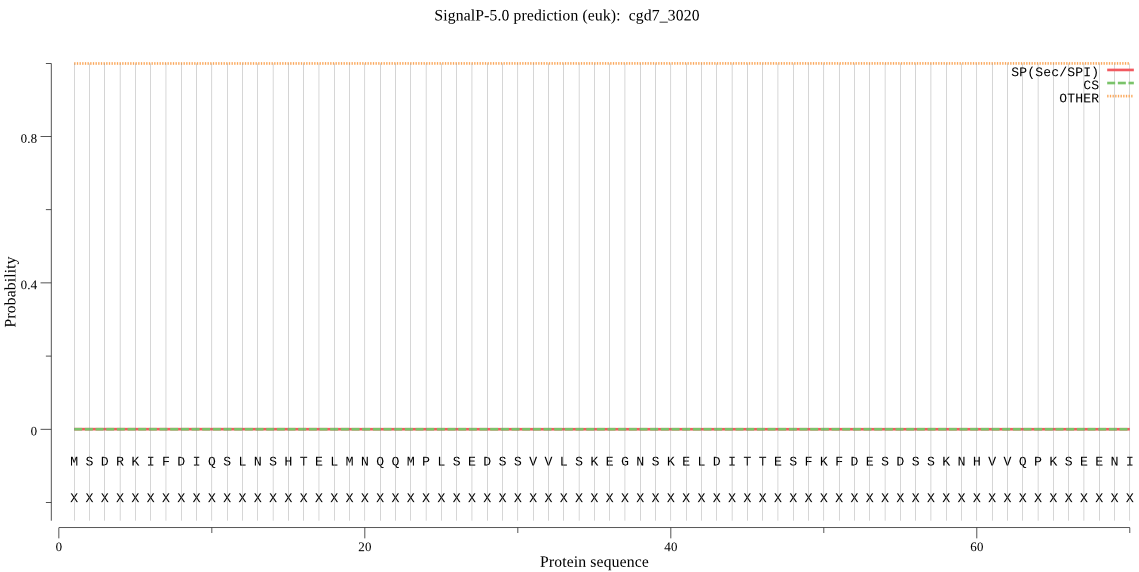
_ID | Prediction | OTHER | SP | mTP | CS_Position |
|
cgd7_3020 | OTHER | 0.999989 | 0.000007 | 0.000004 | |
No Results
-
Fasta :-
>cgd7_3020
MSDRKIFDIQSLNSHTELMNQQMPLSEDSSVVLSKEGNSKELDITTESFKFDESDSSKNH
VVQPKSEENIEGKSSLKNGRTIEIKRGKHTPRNSLFLTMFCCIGGDSSVARLKVSYLATT
FFASSIIFVFIQELVISQLTNGYSIIKIPDGYLFGPPPQVVFDMGALDTNLVRNGQLARL
FWSFWLHTGFIHLFINLSCQIILGIILETRWVIWRYAILYLLGGISGNLASAVLDPCTIS
AGSSACFFALLAGIIVLLLENWRNSRWQFLYVLLVIIASLIGISLSFMSNTDNWAHIGGF
VAGLLWSFASMESFSRKSKALRKSIKSSGKNTIQLPSQDNVQLSLQIKPTDVEVGHSKSQ
FEETSIVDSISDNEAKNEYVKGRFIPANCACSGGVQTVRVISLLLLLSLLTIGFLFLLYK
PLYTKFNLVLGHLSFSGIQKCSCCDSPEGVFWCSSSPTWIRRCG
- Download Fasta
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/cgd7_3020.fa
Sequence name : cgd7_3020
Sequence length : 464
VALUES OF COMPUTED PARAMETERS
Coef20 : 3.859
CoefTot : 0.112
ChDiff : 4
ZoneTo : 2
KR : 0
DE : 0
CleavSite : 0
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 2.482 2.747 0.539 0.912
MesoH : 1.008 1.459 0.139 0.560
MuHd_075 : 29.689 8.945 5.415 4.411
MuHd_095 : 9.906 4.102 2.389 2.484
MuHd_100 : 7.024 4.390 1.735 1.851
MuHd_105 : 14.398 9.440 3.700 3.247
Hmax_075 : 6.300 6.300 -0.530 2.791
Hmax_095 : -1.662 1.488 -2.146 1.234
Hmax_100 : -2.500 3.900 -2.621 1.390
Hmax_105 : 7.000 9.100 0.438 3.372
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.9565 0.0435
DFMC : 0.9462 0.0538
-
Fasta :-
>cgd7_3020
ATGTCTGACAGAAAGATTTTTGATATTCAGTCTTTAAATAGTCATACAGAGTTAATGAAT
CAACAGATGCCCTTAAGTGAAGATAGTTCGGTAGTGCTATCTAAAGAAGGTAATAGTAAG
GAGTTGGATATAACCACAGAATCATTTAAATTTGATGAAAGTGATAGTTCCAAAAATCAT
GTTGTACAACCAAAATCAGAAGAAAATATTGAAGGTAAAAGTTCACTAAAAAATGGAAGA
ACGATTGAAATTAAACGTGGCAAACACACACCTAGGAATTCACTCTTTCTAACGATGTTT
TGTTGTATTGGCGGTGACTCAAGTGTAGCAAGGCTTAAGGTTTCATATTTGGCTACAACC
TTTTTTGCATCTAGTATTATTTTTGTTTTTATACAAGAGCTAGTAATTAGCCAGCTTACG
AACGGTTATTCTATTATAAAAATACCTGATGGATATTTGTTTGGACCTCCTCCTCAGGTA
GTTTTTGATATGGGAGCATTAGATACGAATCTCGTTAGAAATGGGCAGTTGGCTCGACTC
TTTTGGTCATTTTGGCTTCACACTGGATTTATCCATCTCTTCATTAATCTTTCATGTCAA
ATAATACTTGGCATTATTCTAGAGACTCGCTGGGTTATATGGAGATATGCGATACTTTAC
TTATTGGGTGGTATTTCTGGTAATTTAGCCTCTGCAGTCTTGGATCCGTGCACAATCTCT
GCCGGCTCATCAGCTTGCTTCTTTGCGCTTTTGGCAGGGATTATTGTATTACTATTAGAA
AATTGGAGAAATTCTCGGTGGCAATTTTTATACGTTCTATTAGTTATTATTGCCTCTTTA
ATAGGGATTTCCTTAAGTTTCATGTCAAACACAGATAATTGGGCCCATATCGGTGGATTT
GTTGCAGGATTATTATGGTCCTTTGCAAGCATGGAGTCATTTTCAAGAAAATCCAAAGCT
TTACGTAAATCAATAAAATCTTCAGGAAAAAATACGATTCAATTGCCAAGTCAAGATAAT
GTTCAGCTTTCACTTCAAATTAAGCCAACTGATGTTGAAGTCGGTCATTCCAAAAGCCAG
TTTGAAGAAACTTCAATAGTAGATTCTATTTCTGATAACGAAGCTAAAAATGAATATGTT
AAAGGACGCTTTATCCCAGCAAATTGCGCATGCAGCGGCGGTGTTCAAACTGTCAGGGTT
ATCTCTTTACTGTTATTATTATCGTTACTAACTATTGGGTTTTTGTTTTTGTTATATAAG
CCATTGTACACTAAGTTCAATTTGGTTTTAGGCCATCTCTCATTCTCAGGCATACAAAAA
TGCAGTTGCTGCGATTCGCCCGAAGGCGTATTCTGGTGCTCGAGTTCCCCAACATGGATT
AGAAGATGTGGATAA
- Download Fasta
|
ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method |
|
cgd7_3020 | 75 S | EGKSSLKNG | 0.997 | unsp | cgd7_3020 | 75 S | EGKSSLKNG | 0.997 | unsp | cgd7_3020 | 75 S | EGKSSLKNG | 0.997 | unsp | cgd7_3020 | 324 S | ALRKSIKSS | 0.997 | unsp | cgd7_3020 | 328 S | SIKSSGKNT | 0.996 | unsp | cgd7_3020 | 369 S | SIVDSISDN | 0.997 | unsp | cgd7_3020 | 56 S | DESDSSKNH | 0.996 | unsp | cgd7_3020 | 66 S | VQPKSEENI | 0.992 | unsp |


