-
Computed_GO_Component_IDs:
-
Computed_GO_Components:
-
Computed_GO_Function_IDs:
-
Computed_GO_Functions:
-
Computed_GO_Process_IDs:
-
Computed_GO_Processes:
-
Curated_GO_Component_IDs:
-
Curated_GO_Components:
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
No Results
-
Fasta :-
>NF0058120
MLKYTTVWRGLHGVKHKFGTILKSRGVVLSSFGFTNYHHHHDNMIIPHGGSLSLFTSTSL
TTSFRLFSQSSLSRTTLLDSQKVAPEAPQVHSSSNNQKQSSNSNGEQENNKTTHKGFSFL
KFLKKLIKYMTISVASVMGLLAVCLPPALMVDFKEEFENDGYRAVLVALLDAYKDTWNII
ARFTRSCYSIGKVMTIYRNLNKDFEPKFESISRNKRSDHEISSEEELQVLRDYVTTRRQA
NAKAAEILLDLCLHNGGAYVKAGQYIASLNYIMPKEITDILSVLQDKCQQHDFSVTEKIL
LEDLDRDYREIFSSFDTTPIASASIAQVHKATLRDGNQTVAVKIQHPDLIRIFKADLKTM
QFLLFSTKLFFDFPFAWTLPEFERILLSELDFVNEAANCSHFSQMFKDYPDNQIDAPRVH
WNLTSKRILTMDFIEGCKINDLQALKSMNIDPKEVAKLLVDSYSIQTFIHGFVHSDIHVG
NLLVRRSEKNNKPQLIYLDHGCYKKLDEETRKDYCQLWKAAIFRDHENLKLYTEKFGIDG
RFYPLFGLFLTFSNYMDKSSSSMVDQRKNMTSEEAKKMFKQIRETFFPKSTHKAQDLFNI
IEEMFRNMKLDLILLMRANVQLRSITRELGRPINRFAIMAEYCLKGIHYERKNYEMPCIP
HRESDQRRVAVTHVQPTPFFHQLTFDLFKFKLRLLVLDLAFVFFNLSMRILRYFNPSKYE
EAEDDRETSRVKRRLDRMKQQKKAMEEDVASSKEDIEDLSAQQ
- Download Fasta
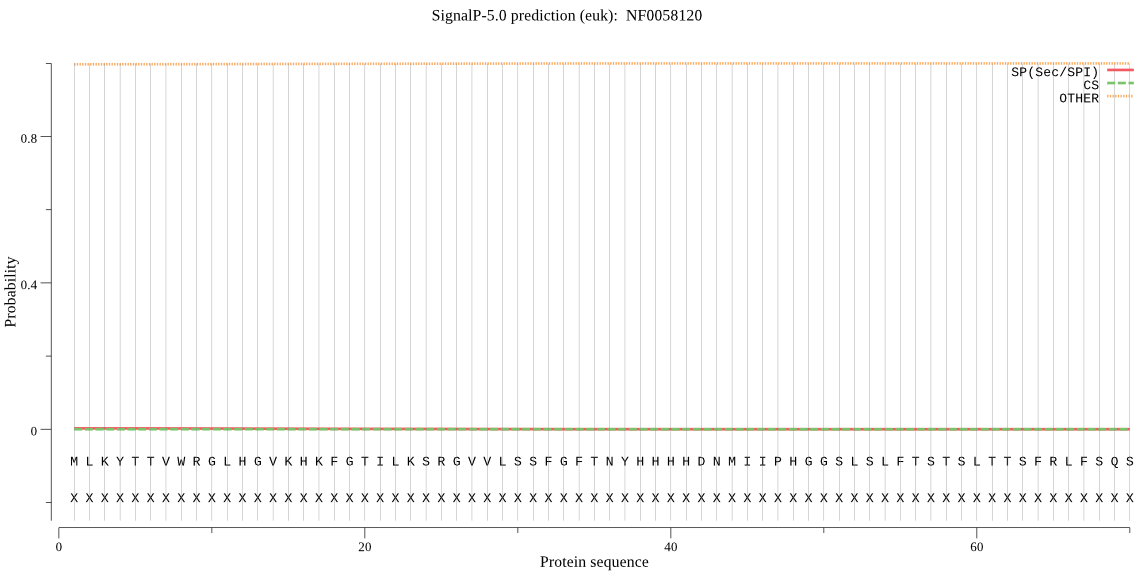
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/NF0058120.fa
Sequence name : NF0058120
Sequence length : 763
VALUES OF COMPUTED PARAMETERS
Coef20 : 3.619
CoefTot : -1.877
ChDiff : 11
ZoneTo : 78
KR : 8
DE : 1
CleavSite : 75
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 2.029 2.065 0.416 0.686
MesoH : -0.233 0.602 -0.320 0.256
MuHd_075 : 34.169 19.292 7.735 7.731
MuHd_095 : 28.229 29.073 10.817 7.896
MuHd_100 : 34.689 32.280 11.900 9.474
MuHd_105 : 32.691 29.115 10.157 8.666
Hmax_075 : 15.100 18.200 2.011 4.220
Hmax_095 : 17.325 22.900 4.246 7.480
Hmax_100 : 18.600 22.900 4.246 7.480
Hmax_105 : 8.100 21.700 3.019 7.828
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.6455 0.3545
DFMC : 0.4339 0.5661
This protein is probably imported in chloroplast.
f(Ser) = 0.1538 f(Arg) = 0.0513 CMi = 1.08499
CMi is the Chloroplast/Mitochondria Index
It has been proposed by Von Heijne et al
(Eur J Biochem,1989, 180: 535-545)
-
Fasta :-
>NF0058120
ATGTTGAAATATACCACTGTTTGGAGAGGACTCCATGGAGTGAAACACAAGTTCGGAACG
ATTCTCAAATCTCGAGGAGTCGTCTTGTCATCTTTTGGATTCACCAATTATCATCATCAT
CATGATAACATGATCATTCCTCATGGAGGTTCTCTCTCGTTATTCACCTCAACCTCACTT
ACCACATCATTCCGACTCTTCTCTCAATCATCCTTGTCTCGAACAACATTATTGGACAGT
CAAAAGGTAGCTCCGGAAGCACCTCAAGTACATTCATCATCGAATAACCAAAAACAGTCA
TCCAATTCGAATGGCGAGCAAGAAAACAACAAAACAACCCACAAGGGGTTTTCCTTTCTT
AAATTCCTCAAAAAGCTCATCAAATACATGACCATTTCAGTTGCCTCTGTTATGGGTTTG
CTTGCGGTGTGTTTACCTCCTGCTTTGATGGTGGATTTCAAAGAGGAGTTTGAAAATGAT
GGTTATAGAGCTGTGCTTGTGGCTCTTTTGGATGCATATAAGGATACGTGGAACATTATC
GCAAGATTTACTCGATCATGCTACTCGATCGGAAAGGTTATGACTATCTATCGAAATCTC
AATAAGGATTTTGAGCCAAAATTTGAAAGTATTTCCAGAAACAAGAGAAGTGACCATGAA
ATTTCTTCGGAAGAGGAACTGCAAGTGTTGCGAGATTATGTGACAACACGAAGACAAGCC
AATGCTAAAGCAGCAGAAATTTTACTTGATCTATGTTTGCACAATGGTGGTGCATATGTG
AAGGCAGGTCAATATATTGCATCGCTCAATTATATTATGCCTAAGGAAATTACAGATATT
CTTTCGGTGCTGCAAGACAAGTGTCAACAACATGATTTTTCAGTCACTGAGAAAATATTA
TTGGAGGACTTGGATCGAGACTATCGAGAAATATTCTCCTCCTTCGATACTACGCCAATA
GCAAGCGCCAGCATTGCACAGGTACATAAAGCCACTTTGAGAGACGGAAATCAAACCGTG
GCTGTCAAAATTCAACATCCAGACTTGATTCGTATTTTTAAGGCTGATTTGAAAACGATG
CAATTTTTACTCTTTTCAACGAAATTATTTTTCGATTTTCCCTTTGCATGGACTTTGCCA
GAGTTTGAGAGAATTTTATTGAGCGAATTAGACTTTGTGAATGAAGCTGCTAATTGTTCA
CATTTTTCTCAAATGTTCAAAGATTACCCTGATAATCAAATTGATGCGCCACGAGTTCAT
TGGAATTTAACAAGCAAACGAATACTAACTATGGATTTTATTGAGGGGTGCAAAATCAAT
GACTTGCAAGCATTGAAAAGCATGAACATTGATCCTAAAGAGGTAGCCAAATTGTTGGTA
GATAGTTATTCCATTCAGACTTTTATTCATGGATTTGTTCATAGTGATATACATGTTGGA
AATTTACTAGTGCGGCGCTCAGAAAAGAATAACAAACCTCAACTCATATATTTAGATCAT
GGATGCTACAAAAAACTCGATGAAGAAACTAGAAAAGATTATTGTCAATTGTGGAAGGCT
GCAATTTTTAGAGATCATGAGAATTTGAAATTATATACCGAAAAATTTGGAATTGACGGA
AGATTTTATCCCCTCTTTGGGTTGTTCTTGACCTTTTCAAATTATATGGATAAAAGTTCA
AGTTCCATGGTGGACCAACGAAAGAACATGACCAGTGAAGAAGCAAAGAAAATGTTTAAA
CAAATTAGAGAGACCTTTTTCCCTAAAAGCACTCATAAGGCACAAGATTTATTCAACATT
ATCGAAGAAATGTTCCGCAATATGAAATTGGATCTTATTTTACTCATGAGAGCCAATGTT
CAATTAAGATCCATCACAAGAGAGTTGGGTCGACCAATCAATCGATTCGCCATCATGGCA
GAATATTGTCTGAAAGGAATTCATTATGAAAGAAAAAATTATGAAATGCCTTGCATTCCA
CATCGAGAATCAGATCAGAGACGAGTCGCTGTCACTCATGTTCAACCAACACCCTTCTTT
CATCAACTCACGTTTGATTTATTCAAATTTAAGCTCCGTTTACTTGTGTTAGATTTGGCA
TTTGTATTTTTCAACTTGTCGATGCGAATATTGAGATATTTCAATCCTTCCAAATATGAG
GAAGCAGAAGATGACAGGGAAACTTCACGAGTGAAAAGAAGATTGGATCGAATGAAACAA
CAAAAGAAAGCTATGGAAGAAGATGTAGCCTCTTCGAAAGAGGACATTGAAGATTTATCG
GCTCAACAGTAA
- Download Fasta
- title: putative ATP binding site
- coordinates: L333,R334,D335,G336,Q338,V340,I350,I352,P410,M431,D432,F433,I434,D476,G480,N481,L483,L498,D499
|
ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method |
|
NF0058120 | 222 S | DHEISSEEE | 0.994 | unsp | NF0058120 | 222 S | DHEISSEEE | 0.994 | unsp | NF0058120 | 222 S | DHEISSEEE | 0.994 | unsp | NF0058120 | 223 S | HEISSEEEL | 0.991 | unsp | NF0058120 | 487 S | LVRRSEKNN | 0.997 | unsp | NF0058120 | 562 S | KSSSSMVDQ | 0.998 | unsp | NF0058120 | 717 S | YFNPSKYEE | 0.991 | unsp | NF0058120 | 728 T | DDRETSRVK | 0.99 | unsp | NF0058120 | 751 S | EDVASSKED | 0.995 | unsp | NF0058120 | 752 S | DVASSKEDI | 0.998 | unsp | NF0058120 | 80 S | TLLDSQKVA | 0.991 | unsp | NF0058120 | 103 S | QSSNSNGEQ | 0.997 | unsp |


