-
Computed_GO_Component_IDs:
-
Computed_GO_Components:
-
Computed_GO_Function_IDs:
-
Computed_GO_Functions:
-
Computed_GO_Process_IDs:
-
Computed_GO_Processes:
-
-
-
Curated_GO_Function_IDs:
-
Curated_GO_Functions:
-
Curated_GO_Processes:
No Results
-
Fasta :-
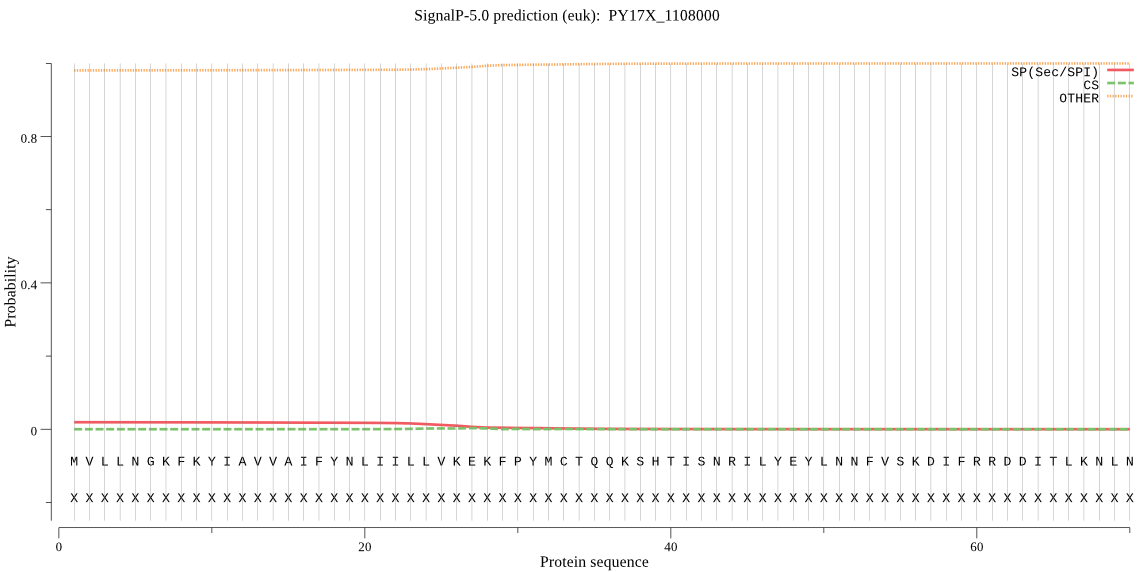
>PY17X_1108000
MVLLNGKFKYIAVVAIFYNLIILLVKEKFPYMCTQQKSHTISNRILYEYLNNFVSKDIFR
RDDITLKNLNFVETNLKNDKDAEIKENSDTQSVDDNMFQRVYKFVLNFFYGNKKNRINKS
MNYGRYDNFNKINDIFEFMRNNGLPINITSVCLIDTGLNIKDELINYFLNHDISTYNSYT
YHSVNINYKKPDSFNLGVNSENCDEDSNSECESTFLENHNGYEKYEDRSKIQGDTLKLIE
KKYEKNVDLQKSGINVEICKTFDNSKEKKNDSNIIPVIKCLEYCKTKNVKIIHIDYNINE
KNEQLMQIIEDLKKSEIFVVLPSKKLFNEKSHGDNSVIYPFPFFEDFEHFEKFEKFENVF
FLDSINSSLKKGDDHDMLDYEIRYSSAFFINIITIILNIYPNISIKELRNILSYSILSKE
TPELKTENIFEGNFDINRFIHILLNRGIIYSDFVQKYEDVTPNEPTNKTSVLEGQEDGEI
EPQTDSSTDIYYDEDGSGEGGSDEGGSGEGGSGEGGSGEGGSGEGGSGEGGSGEGGSGEG
GSGEDVLSTLNELDDYSEDSYSNNKSEQLLEDENGLEDEPLLEDENEFEGETSKDLYSVK
ENDIYVIESKNPTNSSFMQMNYDDRIKSKYVDNLEDANYQNHRAQFEINRNGNRYPIFSE
DNLRDRQNTHNIQMVNDGINYSENGRNIEELYDNDFEGYRGKNLNPEYNNNNNNNNNINR
EFGILNSWRKNDEGLYLNDNEEASQPFEWMDMHADDIYQDRKKRLKENNYDNKNGRGRRI
DEDNKRGRYIKNRNMQKKKKFDRDVKNKRYLNRRRSKRQISKKNKRIMREKKNKYNNSIL
KRNEMKSHNNPQKSPKIIPRRYSR
- Download Fasta
-
MitoProt II - v1.101
File : /home/rajan/sadaf/4480_mitoprot/test/PY17X_1108000.fa
Sequence name : PY17X_1108000
Sequence length : 864
VALUES OF COMPUTED PARAMETERS
Coef20 : 4.169
CoefTot : -1.507
ChDiff : -30
ZoneTo : 47
KR : 6
DE : 1
CleavSite : 0
HYDROPHOBIC SCALE USED
GES KD GVH1 ECS
H17 : 1.312 2.259 0.184 0.764
MesoH : -0.953 0.317 -0.472 0.209
MuHd_075 : 25.839 18.694 8.105 5.398
MuHd_095 : 23.057 20.734 9.539 5.265
MuHd_100 : 19.218 13.933 6.241 4.752
MuHd_105 : 26.082 15.972 8.548 5.827
Hmax_075 : 17.617 18.700 4.803 6.743
Hmax_095 : 7.700 19.162 0.784 2.770
Hmax_100 : 16.900 19.100 4.236 7.280
Hmax_105 : 17.200 23.217 4.236 7.560
CLASS NOT-MITO MITO(/CHLORO)
DFM : 0.9208 0.0792
DFMC : 0.9302 0.0698
-
Fasta :-
>PY17X_1108000
ATGGTCCTATTAAATGGAAAATTCAAATATATCGCGGTTGTAGCGATATTTTACAATCTT
ATTATTTTATTAGTTAAGGAAAAGTTTCCTTATATGTGCACCCAACAAAAATCCCATACT
ATTAGTAATAGAATTCTTTATGAATATTTGAATAATTTTGTATCCAAAGATATTTTTAGA
CGCGACGATATAACTCTTAAGAATCTTAACTTTGTTGAAACAAACTTGAAAAATGATAAG
GATGCTGAAATTAAAGAAAATAGTGATACACAAAGCGTTGATGATAATATGTTTCAAAGG
GTGTATAAATTTGTTTTGAATTTTTTTTATGGAAATAAGAAAAATCGTATAAACAAATCC
ATGAACTATGGAAGATATGATAATTTTAATAAAATAAATGATATTTTTGAATTTATGAGA
AACAATGGTTTACCAATTAATATAACATCAGTATGCTTAATAGACACTGGATTAAATATT
AAGGATGAGTTAATTAACTATTTTTTAAATCATGATATTAGCACATATAATAGTTATACA
TATCATTCAGTTAATATAAATTATAAAAAACCTGATTCATTTAATTTGGGAGTAAATAGC
GAAAATTGTGATGAAGATAGTAATTCTGAATGTGAGTCTACATTTTTGGAGAATCATAAT
GGGTATGAAAAATATGAAGATAGATCAAAAATCCAAGGCGATACATTAAAATTGATAGAA
AAAAAATATGAAAAAAATGTGGATTTACAAAAGAGTGGTATAAATGTAGAAATATGCAAA
ACATTTGATAATAGTAAGGAAAAAAAAAATGATTCGAATATAATTCCTGTAATTAAATGT
CTAGAATATTGTAAAACAAAAAATGTAAAAATAATTCATATAGATTATAATATAAATGAA
AAAAATGAACAATTAATGCAAATTATAGAAGATTTAAAAAAATCGGAAATATTCGTAGTT
TTACCATCGAAAAAACTATTTAATGAAAAGTCTCATGGAGACAATTCTGTAATATATCCA
TTTCCATTTTTTGAAGATTTTGAACATTTTGAAAAATTTGAAAAATTTGAAAATGTTTTT
TTTTTGGACAGTATAAATAGTAGTTTAAAAAAAGGAGATGACCATGATATGTTAGATTAT
GAAATTAGATATAGTTCAGCATTTTTTATAAATATAATAACAATCATTCTAAATATATAT
CCCAATATATCTATTAAAGAGTTACGAAATATATTAAGCTATTCAATACTATCGAAAGAA
ACGCCAGAATTAAAAACAGAAAATATTTTTGAAGGTAACTTTGACATAAATAGATTTATT
CATATTTTATTAAATAGGGGAATAATTTACTCTGATTTTGTTCAAAAATATGAAGATGTA
ACACCGAATGAGCCCACGAATAAAACATCTGTTTTAGAGGGTCAGGAAGATGGTGAGATA
GAACCACAGACCGATTCCTCTACTGATATATACTATGATGAAGATGGTTCTGGTGAAGGT
GGTTCTGATGAAGGTGGTTCTGGTGAAGGTGGTTCTGGTGAAGGTGGTTCTGGTGAAGGT
GGTTCTGGTGAAGGTGGTTCTGGTGAAGGTGGTTCTGGTGAAGGTGGTTCTGGTGAAGGT
GGTTCTGGTGAAGATGTATTGAGTACTTTGAATGAGTTGGATGACTATAGTGAGGATTCC
TATAGCAATAATAAGAGTGAGCAGTTATTAGAGGATGAAAATGGGCTTGAAGATGAGCCG
TTATTAGAGGATGAAAATGAATTTGAAGGTGAGACTTCCAAAGATTTATATAGTGTTAAA
GAAAACGATATTTATGTTATTGAAAGTAAGAATCCAACTAATAGCTCTTTTATGCAAATG
AATTATGATGATAGAATAAAATCAAAATATGTAGACAATTTAGAAGATGCAAATTATCAA
AATCATAGAGCTCAATTTGAGATAAATAGAAATGGCAATAGATATCCTATATTTTCTGAA
GACAATTTAAGAGATAGACAAAATACGCATAACATTCAGATGGTAAACGATGGTATAAAT
TATTCAGAAAATGGAAGGAATATAGAAGAATTATATGATAATGATTTTGAAGGTTATAGA
GGAAAAAATTTAAACCCCGAGTATAATAATAACAATAATAATAATAATAATATAAATAGA
GAATTCGGAATTTTAAATTCTTGGAGAAAAAATGATGAGGGATTATATTTGAATGATAAT
GAAGAAGCTAGTCAGCCATTTGAATGGATGGACATGCATGCAGATGATATATATCAAGAT
CGAAAAAAACGATTAAAGGAAAATAATTATGATAATAAGAATGGAAGAGGAAGAAGAATA
GATGAAGATAATAAACGAGGTAGATATATAAAGAATCGAAATATGCAGAAAAAAAAAAAA
TTTGATCGTGATGTAAAAAATAAAAGATATTTAAATAGAAGAAGGAGTAAAAGACAGATC
AGTAAGAAAAATAAAAGGATTATGAGAGAAAAAAAAAATAAATATAATAATAGTATACTA
AAAAGAAATGAAATGAAATCACATAATAATCCACAAAAAAGTCCCAAAATAATTCCAAGA
AGATATTCTAGATAA
- Download Fasta
|
ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method | ID | Site | Peptide | Score | Method |
|
PY17X_1108000 | 560 S | YSEDSYSNN | 0.993 | unsp | PY17X_1108000 | 560 S | YSEDSYSNN | 0.993 | unsp | PY17X_1108000 | 560 S | YSEDSYSNN | 0.993 | unsp | PY17X_1108000 | 561 Y | SEDSYSNNK | 0.994 | unsp | PY17X_1108000 | 598 S | KDLYSVKEN | 0.998 | unsp | PY17X_1108000 | 816 S | NRRRSKRQI | 0.997 | unsp | PY17X_1108000 | 821 S | KRQISKKNK | 0.998 | unsp | PY17X_1108000 | 854 S | NPQKSPKII | 0.991 | unsp | PY17X_1108000 | 209 S | EDSNSECES | 0.994 | unsp | PY17X_1108000 | 404 S | YPNISIKEL | 0.996 | unsp |


