Viruses are among the most agriculturally important group of plant pathogens, causing serious economic losses in many major crops by reducing yield and quality. Many researches have shown that plants use RNA interference as an antiviral defence mechanism. During replication of viruses at a given point of their life cycle, they reach the stage of ssRNA or dsRNA which can trigger host defence response.
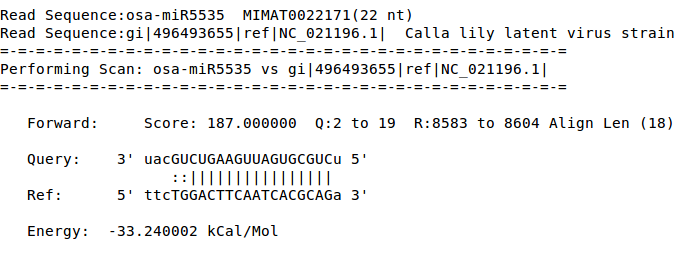
In current study interaction between plant viral genome and plant miRNA are compiled as a database. miRNA from seven different species (A.thaliana, G.max, O.sativa, S.bicolor, S.licopersicum, V.vinifera, Z.mays) and approx 500 viral genomes majorly from geminivirus family and potyvirus family are taken into account. Targets for each set of miRNAs were searched in the viral genome using a modified version of miRanda.
Researches in last few years gave evidence about the indispensable role of miRNA in antiviral activities in plants, but in bioinformatic approaches alignment details were missing and in experimental evidence problem is confined to a few miRNAs only, moreover there in no Plant database available which have a collection of potential miRNA targets in the viral genome. This is first ever database which encompasses 7 major plant species and 500 viral species interactions. The interface of Database gives an opportunity to select results based on free energy threshold, Query cover, identity percentage and interaction site interaction site restriction etc which make it more useful for researchers to get most probable targets in viral genomes. Results contain complete detail about alignment also which makes the results more useful for further investigation. This could be prove as must visit destination for every beginner in Plant miRNA and viral interaction domain.
This database can be seen as the collection for the potential targets of plant miRNAs in viral genomes, which are likely to have played a role in early plant evolution, shaping host range for plant infecting viruses and having antiviral activities.