PAmiRDB is a database of computationally predicted whole genome viral targets of plant miRNA.
The database comprises of predicted and preferred interactions between Plant virus targets (The majority of viruses belongs to Geminiviridae and potyviridae) with 2842
miRNA(updated according to miRbase release 22- march 2018) from six plant species (A. thaliana, G. max, O. sativa, S. bicolor, V. vinifera, Z. mays, S. lycopersicum).
PAmiRDb is updated with every release of miRBase so that no relevent miRNA left outside the purview of the database.
The user friendly interface of PAmiRDB provides intuitive interface to search the database using GenomeID of virus, miRNAID, Virus name, Binding Energy, and Target proteins. The
query may also be customised by restriction of different specific binding positions.
The general miRNA-target binding conditions which are taken into consideration are as follows:
1. four or fewer mismatches in overall alignment.
2. No mismatch in seed regions (position 2-8).
3. No more than two consecutive mismatches in position 13-21.
4. Max energy -20(kCal/Mol).
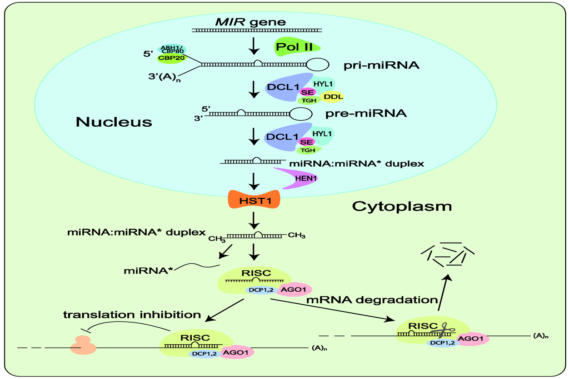
The research conducted by various groups indicate that miRNAs can guide transcriptional silencing in plants. miRNAs negatively regulate gene expression by targeting specific mRNA targets for cleavage or translational inhibition. miRNAs are an important component of plants' response to various biotic and abiotic stresses including infection by the viral pathogen. RNA silencing is a eukaryotic surveillance mechanism against invasive nucleic acid including plant viruses. With the rapidly developing interest in plant miRNA and viral genome interactions amongst research community, we anticipate that the database will be a useful to the researchers. PAmiRDB is developed and maintained by Translational Bioinformatics Group, ICGEB New Delhi India.